Key Documents
I7634
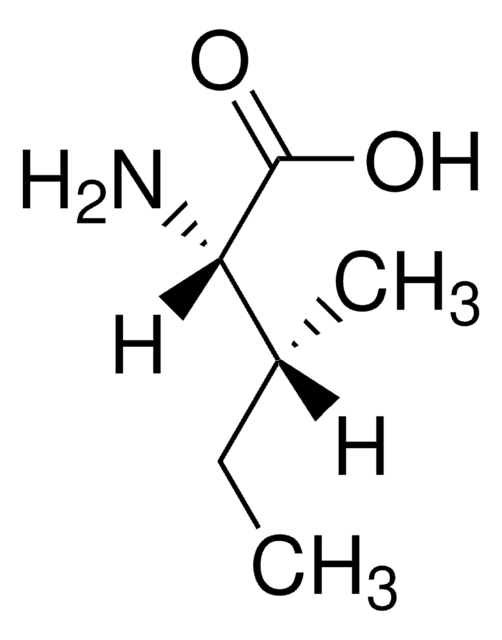
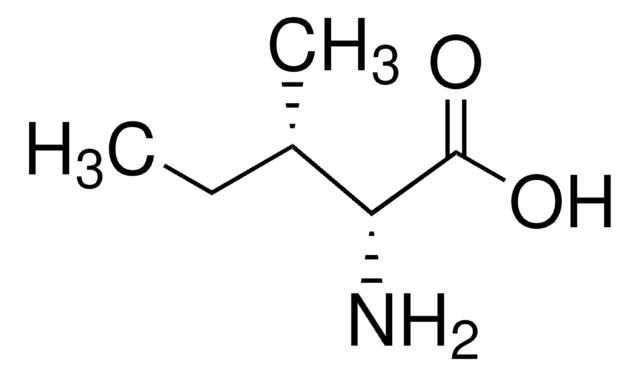
D-Isoleucine
≥98% (TLC)
Synonim(y):
(2R, 3R)-2-Amino-3-methylpentanoic acid
About This Item
Polecane produkty
product name
D-Isoleucine, ≥98% (TLC)
Próba
≥98% (TLC)
Postać
powder
zanieczyszczenia
≤10% allo-isomer
kolor
white
Zastosowanie
cell analysis
ciąg SMILES
CC[C@@H](C)[C@@H](N)C(O)=O
InChI
1S/C6H13NO2/c1-3-4(2)5(7)6(8)9/h4-5H,3,7H2,1-2H3,(H,8,9)/t4-,5-/m1/s1
Klucz InChI
AGPKZVBTJJNPAG-RFZPGFLSSA-N
Szukasz podobnych produktów? Odwiedź Przewodnik dotyczący porównywania produktów
Powiązane kategorie
Zastosowanie
- Metabolic Profiling of Urinary Chiral Amino-Containing Biomarkers for Gastric Cancer Using a Sensitive Chiral Chlorine-Labeled Probe by HPLC-MS/MS.: This study by Huang et al. (2021) explores the use of D-Isoleucine as a biomarker for gastric cancer, utilizing advanced metabolic profiling techniques with HPLC-MS/MS to detect chiral amino-containing compounds in urine samples (Huang et al., 2021).
- Isopenicillin N Synthase Binds δ-(L-α-aminoadipoyl)-L-cysteinyl-D-thia-allo-isoleucine Through Both Sulfur Atoms.: Clifton et al. (2011) describe the binding mechanism of Isopenicillin N synthase with a compound involving D-Isoleucine, highlighting its role in antibiotic biosynthesis and enzyme interaction studies (Clifton et al., 2011).
- The Total Structure of the Antibiotic Longicatenamycin.: Shiba and Mukunoki (1975) determined the complete structure of the antibiotic longicatenamycin, which includes D-Isoleucine as a crucial component, contributing to understanding its antibacterial properties (Shiba and Mukunoki, 1975).
Działania biochem./fizjol.
Zastosowanie
Kod klasy składowania
11 - Combustible Solids
Klasa zagrożenia wodnego (WGK)
WGK 3
Temperatura zapłonu (°F)
Not applicable
Temperatura zapłonu (°C)
Not applicable
Certyfikaty analizy (CoA)
Poszukaj Certyfikaty analizy (CoA), wpisując numer partii/serii produktów. Numery serii i partii można znaleźć na etykiecie produktu po słowach „seria” lub „partia”.
Masz już ten produkt?
Dokumenty związane z niedawno zakupionymi produktami zostały zamieszczone w Bibliotece dokumentów.
Klienci oglądali również te produkty
Chromatograms
application for HPLCNasz zespół naukowców ma doświadczenie we wszystkich obszarach badań, w tym w naukach przyrodniczych, materiałoznawstwie, syntezie chemicznej, chromatografii, analityce i wielu innych dziedzinach.
Skontaktuj się z zespołem ds. pomocy technicznej