Wszystkie zdjęcia(3)
Key Documents
908401
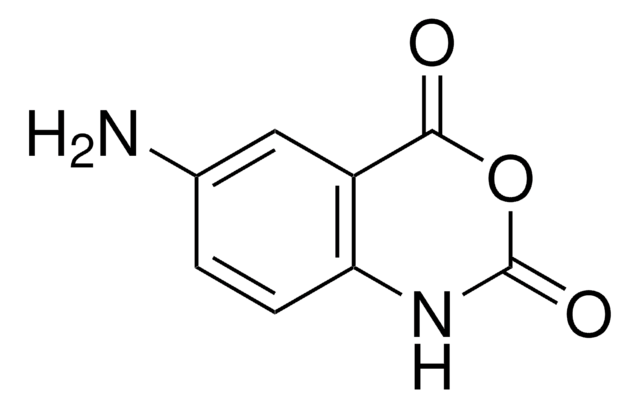
1-Methyl-7-nitroisatoic anhydride
Synonim(y):
1-Methyl-7-nitro-2H-3,1-benzoxazine-2,4(1H)-dione, 1-methyl-7-nitro-2H-3,1-Benzoxazine-2,4(1H)-dione, 1M7, RNA SHAPE probe, Reagent for RNA SHAPE-MaP
Zaloguj sięWyświetlanie cen organizacyjnych i kontraktowych
About This Item
Wzór empiryczny (zapis Hilla):
C9H6N2O5
Numer CAS:
Masa cząsteczkowa:
222.15
Numer MDL:
Kod UNSPSC:
12352119
NACRES:
NA.22
Polecane produkty
Postać
powder
mp
204.5 °C
temp. przechowywania
2-8°C
InChI
1S/C9H6N2O5/c1-10-7-4-5(11(14)15)2-3-6(7)8(12)16-9(10)13/h2-4H,1H3
Klucz InChI
MULNCJWAVSDEKJ-UHFFFAOYSA-N
Zastosowanie
1-Methyl-7-nitroisatoic anhydride (1M7) is used as an in vivo SHAPE-MaP reagent for live cell RNA structure analysis at single nucleotide resolution. SHAPE -- or selective 2′-hydroxyl acylation analyzed by primer extension -- uses small, electrophilic chemical probes such as 1M7 to react with the 2′-hydroxyl group and provides insight to RNA structure. When combined with mutational profiling (MaP), quantitative nucleotide measurements are possible for entire transciptomes. Together, these methods deepen the understanding of RNA interactions and regions that may be exploited for design of RNA-targeting small-molecule drugs.
Inne uwagi
Pervasive Regulatory Functions of mRNA Structure Revealed by High-Resolution SHAPE Probing
SnapShot: RNA Structure Probing Technologies
Detection of RNA-Protein Interactions in Living Cells with SHAPE
Standardization of RNA Chemical Mapping Experiments
In-cell RNA structure probing with SHAPE-MaP
A Fast-Acting Reagent for Accurate Analysis of RNA Secondary and Tertiary Structure by SHAPE Chemistry
SnapShot: RNA Structure Probing Technologies
Detection of RNA-Protein Interactions in Living Cells with SHAPE
Standardization of RNA Chemical Mapping Experiments
In-cell RNA structure probing with SHAPE-MaP
A Fast-Acting Reagent for Accurate Analysis of RNA Secondary and Tertiary Structure by SHAPE Chemistry
This page may contain text that has been machine translated.
produkt powiązany
Numer produktu
Opis
Cennik
Kod klasy składowania
11 - Combustible Solids
Klasa zagrożenia wodnego (WGK)
WGK 3
Temperatura zapłonu (°F)
Not applicable
Temperatura zapłonu (°C)
Not applicable
Wybierz jedną z najnowszych wersji:
Masz już ten produkt?
Dokumenty związane z niedawno zakupionymi produktami zostały zamieszczone w Bibliotece dokumentów.
Katherine E Deigan et al.
Proceedings of the National Academy of Sciences of the United States of America, 106(1), 97-102 (2008-12-26)
Almost all RNAs can fold to form extensive base-paired secondary structures. Many of these structures then modulate numerous fundamental elements of gene expression. Deducing these structure-function relationships requires that it be possible to predict RNA secondary structures accurately. However, RNA
Scott P Hennelly et al.
Nucleic acids research, 39(6), 2416-2431 (2010-11-26)
Riboswitches are non-coding RNAs that control gene expression by sensing small molecules through changes in secondary structure. While secondary structure and ligand interactions are thought to control switching, the exact mechanism of control is unknown. Using a novel two-piece assay
Matthew J Smola et al.
Biochemistry, 54(46), 6867-6875 (2015-11-07)
SHAPE-MaP is unique among RNA structure probing strategies in that it both measures flexibility at single-nucleotide resolution and quantifies the uncertainties in these measurements. We report a straightforward analytical framework that incorporates these uncertainties to allow detection of RNA structural
Anthony M Mustoe et al.
Cell, 173(1), 181-195 (2018-03-20)
mRNAs can fold into complex structures that regulate gene expression. Resolving such structures de novo has remained challenging and has limited our understanding of the prevalence and functions of mRNA structure. We use SHAPE-MaP experiments in living E. coli cells to
Jacob K Grohman et al.
Journal of the American Chemical Society, 133(50), 20326-20334 (2011-12-01)
Higher-order structure influences critical functions in nearly all noncoding and coding RNAs. Most single-nucleotide resolution RNA structure determination technologies cannot be used to analyze RNA from scarce biological samples, like viral genomes. To make quantitative RNA structure analysis applicable to
Nasz zespół naukowców ma doświadczenie we wszystkich obszarach badań, w tym w naukach przyrodniczych, materiałoznawstwie, syntezie chemicznej, chromatografii, analityce i wielu innych dziedzinach.
Skontaktuj się z zespołem ds. pomocy technicznej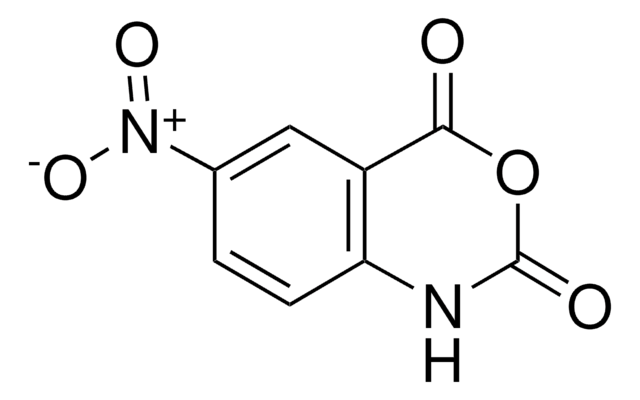
![1,4-Diazabicyclo[2.2.2]octane ReagentPlus®, ≥99%](/deepweb/assets/sigmaaldrich/product/structures/366/129/a6ff4175-974d-4fac-9038-b35e508ef252/640/a6ff4175-974d-4fac-9038-b35e508ef252.png)
![1,8-Diazabicyclo[5.4.0]undec-7-ene 98%](/deepweb/assets/sigmaaldrich/product/structures/120/564/5b373e23-1624-489c-8efb-692de0f96ffb/640/5b373e23-1624-489c-8efb-692de0f96ffb.png)