12158167001
Roche
Anti-HA-Biotin, High Affinity (3F10)
from rat IgG1
Synonym(s):
antibody
Sign Into View Organizational & Contract Pricing
All Photos(1)
About This Item
UNSPSC Code:
12352203
Recommended Products
biological source
rat
Quality Level
conjugate
biotin conjugate
antibody form
purified immunoglobulin
antibody product type
primary antibodies
clone
3F10, monoclonal
form
lyophilized (stabilized)
packaging
pkg of 50 μg
manufacturer/tradename
Roche
isotype
IgG1
epitope sequence
YPYDVPDYA
storage temp.
2-8°C
Related Categories
General description
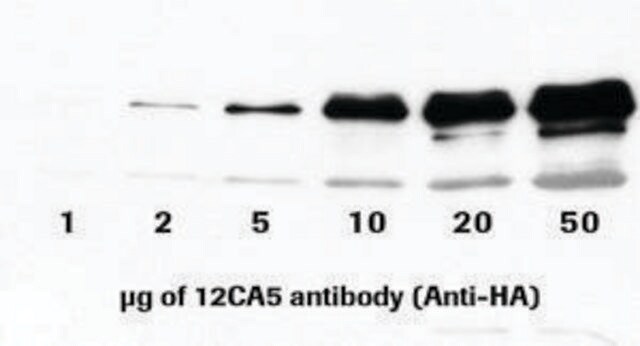
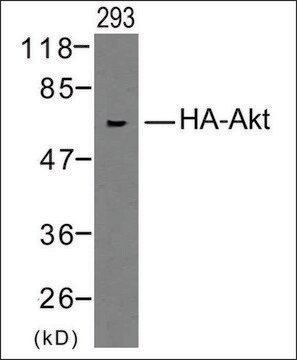
Anti-HA-Biotin, High Affinity (3F10) is a monoclonal antibody for the highly sensitive detection of HA-tagged recombinant proteins, Fab fragments, conjugated to biotin. The Anti-HA-Biotin, High Affinity antibody (clone 3F10) recognizes the same epitope as clone 12CA5. It is a monoclonal antibody whose high affinity and low working concentrations result in less cross-reactivity than with other antibodies to the HA-epitope. Anti-HA-Biotin, High Affinity (3F10) is a biotin conjugate of this clone which is specifically useful in western blotting, ELISA applications and assays using the universal biotin-streptavidin platform, by allowing specific and highly sensitive detection of HA-tagged proteins.
Specificity
Anti-HA-Biotin, High Affinity (3F10) recognizes the 9-amino acid sequence YPYDVPDYA, derived from the human influenza hemagglutinin (HA) protein. This epitope is also recognized in fusion proteins regardless of its position (N-terminal, C-terminal or internal).
Immunogen
Amino acids 98-106 from the human influenza virus hemagglutinin protein
Application
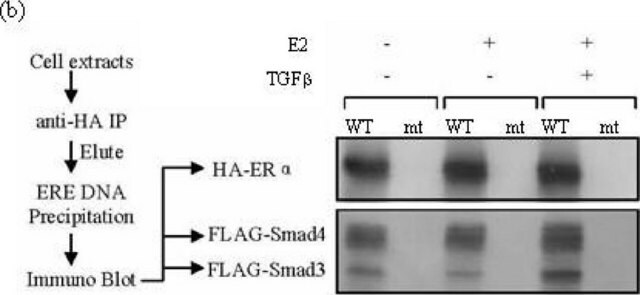
Anti-HA-Biotin, High Affinity (3F10) is used for the detection of HA-tagged recombinant proteins using:
It has also been used for immunocytochemistry, immunofluorescence and αScreen format based assay.
- Dot blots
- ELISA (enzyme-linked immunosorbent assay)
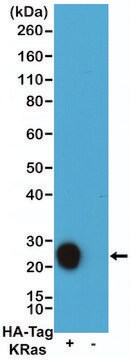
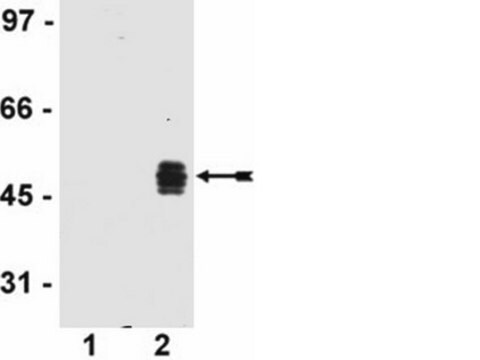
- Western blots
It has also been used for immunocytochemistry, immunofluorescence and αScreen format based assay.
Quality
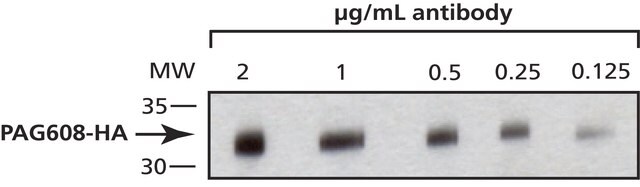
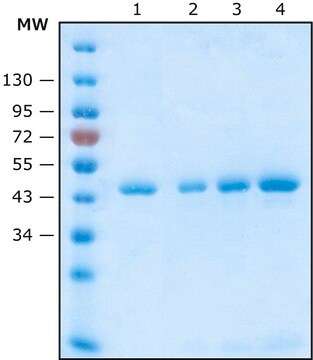
Function test: The Anti-HA-Biotin; High Affinity is function tested by Western blot analysis of a HA-tagged fusion protein.
Preparation Note
Sample Materials
Sample preparation: Prepare protein extracts containing the HA-tagged protein of interest using any of a variety of standard methods. The following lysis buffers have performed well and should be taken as guidelines:
Sample preparation: Prepare protein extracts containing the HA-tagged protein of interest using any of a variety of standard methods. The following lysis buffers have performed well and should be taken as guidelines:
- Bacterial extracts: 20 mM Tris, pH 8.0, 100 mM NaCl, cOmplete Protease Inhibitor Cocktail Tablets, followed by freeze-thaw.
- Mammalian extracts: 50 mM Tris, pH 7.5, 150 mM NaCl, 0.1% Nonidet P40, complete Protease Inhibitor Cocktail Tablets.
- Other cell lysis buffers may be more appropriate for individual applications. In general, to obtain optimal performance of the affinity matrix:
- Use protease inhibitors to reduce proteolytic activity. Use complete Protease Inhibitor Cocktail Tablets for most applications.
- Limit detergent to the lowest concentration levels necessary to obtain adequate cell lysis.
Working concentration: Working concentration of conjugate depends on application and substrate
The following concentrations should be taken as a guideline:
The following concentrations should be taken as a guideline:
- Dot blot: 100 ng/ml
- ELISA: 100 ng/ml
- Western blot: 100 ng/ml
Reconstitution
Add 1 ml double-distilled water to a final concentration of 50 μg/ml.
Rehydrate for 10 minutes prior to use.
Rehydrate for 10 minutes prior to use.
Other Notes
For life science research only. Not for use in diagnostic procedures.
Not finding the right product?
Try our Product Selector Tool.
signalword
Warning
hcodes
Hazard Classifications
Aquatic Chronic 3 - Skin Sens. 1
Storage Class
11 - Combustible Solids
wgk_germany
WGK 2
flash_point_f
does not flash
flash_point_c
does not flash
Choose from one of the most recent versions:
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
Edward J Hsieh et al.
Archives of biochemistry and biophysics, 463(1), 19-26 (2007-03-30)
Coenzyme Q (Q) is a redox active lipid that is an essential component of the electron transport chain. Here, we show that steady state levels of Coq3, Coq4, Coq6, Coq7 and Coq9 polypeptides in yeast mitochondria are dependent on the
Paul A Rowley et al.
PLoS pathogens, 12(10), e1005890-e1005890 (2016-10-07)
In eukaryotes, the degradation of cellular mRNAs is accomplished by Xrn1 and the cytoplasmic exosome. Because viral RNAs often lack canonical caps or poly-A tails, they can also be vulnerable to degradation by these host exonucleases. Yeast lack sophisticated mechanisms
Derek C Prosser et al.
Journal of cell science, 128(22), 4220-4234 (2015-10-16)
Clathrin-mediated endocytosis (CME) is a well-studied mechanism to internalize plasma membrane proteins; however, to endocytose such cargo, most eukaryotic cells also use alternative clathrin-independent endocytic (CIE) pathways, which are less well characterized. The budding yeast Saccharomyces cerevisiae, a widely used
Kenji Tamura et al.
Oncology letters, 14(6), 6650-6658 (2018-01-19)
The present study aimed at identifying novel molecular cancer drug targets and biomarkers by analyzing the gene expression profiles of high-grade prostate cancer (PC), using a cDNA microarray combined with laser microbeam microdissection. A number of genes were identified that
Matthew D Marsden et al.
PLoS pathogens, 13(9), e1006575-e1006575 (2017-09-22)
The ability of HIV to establish a long-lived latent infection within resting CD4+ T cells leads to persistence and episodic resupply of the virus in patients treated with antiretroviral therapy (ART), thereby preventing eradication of the disease. Protein kinase C
Global Trade Item Number
| SKU | GTIN |
|---|---|
| 12158167001 | 4061838718518 |
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service