11745808910
Roche
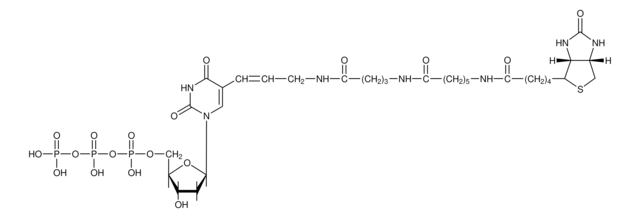
Nick Translation Mix
sufficient for 50 labeling reactions, pkg of 200 μL, solution
Sinônimo(s):
nick translation
About This Item
Produtos recomendados
forma
solution
Nível de qualidade
uso
sufficient for 50 labeling reactions
embalagem
pkg of 200 μL
fabricante/nome comercial
Roche
técnica(s)
nucleic acid labeling: suitable
cor
colorless
pH
~7.5 (68 °F)
solubilidade
water: miscible
adequação
suitable for fluorescent labeling techniques
suitable for molecular biology
aplicação(ões)
genomic analysis
life science and biopharma
temperatura de armazenamento
−20°C
Descrição geral
Individual templates produce consistent results in the standard 90-minutes reaction, and result in an average probe length of 200 base pairs up to 500 base pairs.
Assay Time: 100 minutes
Sample Materials
- Supercoiled and linearized plasmid DNA
- Supercoiled and linearized cosmid DNA
- Purified PCR products
Especificidade
Aplicação
The Nick Translation Mix is designed for direct fluorophore-labeling of in situ probes. Fluorescein-12-dUTP and Tetramethyl-Rhodamine-5-dUTP from Roche Applied Science or other commercially available fluorophor-labeled nucleotides can be combined with the Nick Translation Mix. Direct fluorophore-labeled in situ probes are used for the detection of multi copy or very large hybridization targets on metaphase chromsomes or interphase nuclei.
For a standard labeling reaction using 1 μg template in 20 μl total reaction volume, 4 μl of 5x concentrated fluorophore labeling mix are required.
Componentes
1 vial with 5x concentrated solution, stabilized reaction buffer in 50% glycerol (v/v), DNA Polymerase I and DNase I.
Qualidade
Princípio
E. coli DNA Polymerase I synthesizes DNA complementary to the intact strand in a 5′?3′ direction using the 3′-OH termini of the nick as a primer. The 5′?3′ exonucleolytic activity of DNA polymerase I simultaneously removes nucleotides in the direction of synthesis. The polymerase activity sequentially replaces the removed nucleotides with isotope-labeled or hapten-labeled deoxyribonucleoside triphosphates. At low temperature (+15°C), the unlabeled DNA in the reaction is thus replaced by newly synthesized labeled DNA.
Armazenamento e estabilidade
Outras notas
Denaturing of the template before nick translation is not required.
Código de classe de armazenamento
12 - Non Combustible Liquids
Classe de risco de água (WGK)
WGK 1
Ponto de fulgor (°F)
does not flash
Ponto de fulgor (°C)
does not flash
Certificados de análise (COA)
Busque Certificados de análise (COA) digitando o Número do Lote do produto. Os números de lote e remessa podem ser encontrados no rótulo de um produto após a palavra “Lot” ou “Batch”.
Já possui este produto?
Encontre a documentação dos produtos que você adquiriu recentemente na biblioteca de documentos.
Os clientes também visualizaram
Nossa equipe de cientistas tem experiência em todas as áreas de pesquisa, incluindo Life Sciences, ciência de materiais, síntese química, cromatografia, química analítica e muitas outras.
Entre em contato com a assistência técnica