CAS9D10AP
CRISPR Cas9-D10A Nickase Plasmid
Synonym(s):
CAS9D10A Plasmid
About This Item
Recommended Products
recombinant
expressed in E. coli
Quality Level
packaging
vial of 50 μL
concentration
20 ng/μL in TE buffer; DNA (1μg of plasmid DNA)
application(s)
CRISPR
promoter
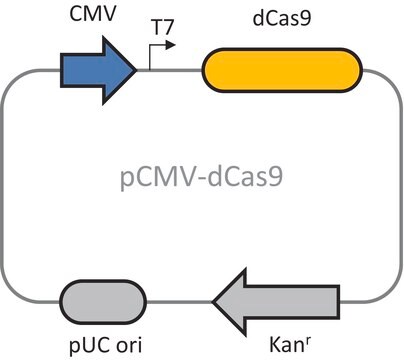
Promoter name: CMV
selection
kanamycin
shipped in
dry ice
storage temp.
−20°C
Looking for similar products? Visit Product Comparison Guide
Related Categories
General description
Application
- Creation of gene knockouts in multiple cell lines
- Complete knockout of genes not amenable to RNAi
- Creation of knock-in cell lines with promoters, fusion tags or reporters integrated into endogenous genes
Features and Benefits
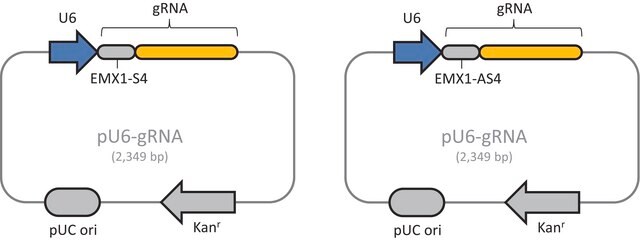
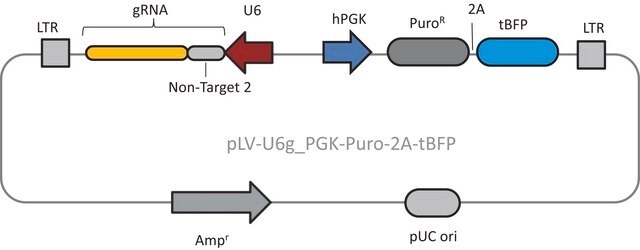
Components
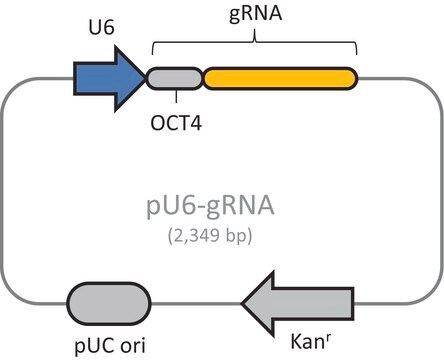
Please note, product does not contain guideRNA sequence. This must be purchased separately through the Custom CRISPR product tab.
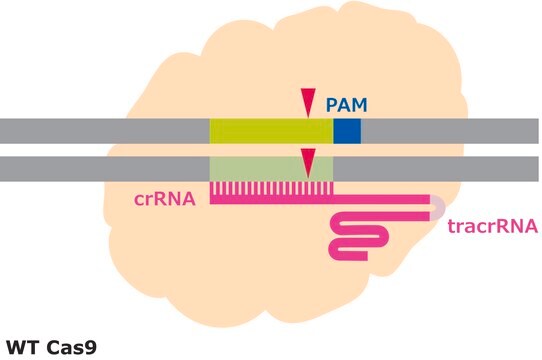
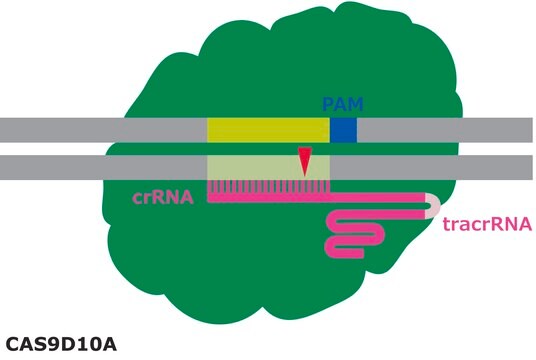
Principle
Physical form
Preparation Note
Other Notes
Typical transfection concentrations used in literature are in the ranges of >= 1.0 ug/uL and <= 5 uL of Cas9-D10A plasmids combined with >= 1.0 ug/uL and <= 5 uL of U6-gRNA plasmids. (All dosages above assume 0.5 to 1 million cells nucleofected)
Legal Information
Storage Class
12 - Non Combustible Liquids
wgk_germany
WGK 1
flash_point_f
Not applicable
flash_point_c
Not applicable
Choose from one of the most recent versions:
Certificates of Analysis (COA)
Don't see the Right Version?
If you require a particular version, you can look up a specific certificate by the Lot or Batch number.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
Articles
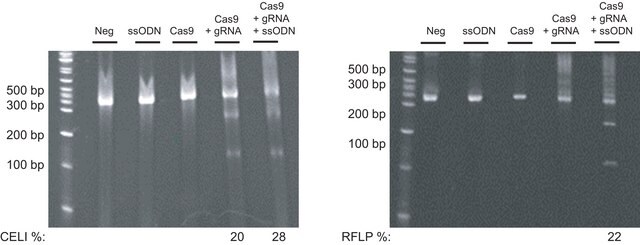
CRISPR endonucleases have shown wide variation in their activity, even among multiple CRISPRs designed within close genomic proximity.
Protocols
Learn about CRISPR Cas9, what it is and how it works. CRISPR is a new, affordable genome editing tool enabling access to genome editing for all.
Related Content
Sigma-Aldrich® Advanced Genomics is the leading provider of gene editing and silencing technologies including CRISPR, Cas9, synthetic guide RNA (sgRNA), and Zinc Finger Nuclease (ZFN).
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service