NOTI-RO
Roche
Not I
from Nocardia otitidis-caviarum
About This Item
Polecane produkty
pochodzenie biologiczne
bacterial (Nocardia otitidis-caviarum)
Poziom jakości
Postać
solution
opakowanie
pkg of 1,000 U (11014714001 [10 U/μl])
pkg of 1,000 U (11037668001 [40 U/μl])
pkg of 200 U (11014706001 [10 U/μl])
producent / nazwa handlowa
Roche
Parametry
37 °C optimum reaction temp.
metody
PCR: suitable
temp. przechowywania
−20°C
Powiązane kategorie
Opis ogólny
Not I belongs to the class of "rare-cutter" enzymes. It is one of the two known enzymes that recognize an octameric sequence comprised solely of G and C residues.
Contents:
- Not I
- SuRE/Cut Buffer H (10x)
Specyficzność
GCGG*C*CG °C
Restriction site: GC↓GG*C*CG °C
GC↓GG*C*CG °C
Heat inactivation: Not I can be heat inactivated by incubation at 65 °C for 15 minutes (up to 100 U/μg DNA).
Jakość
1 μg Ad2 DNA is incubated for 16 hours in 50 μl SuRE/Cut Buffer H with an excess of Not I. The number of enzyme units which do not change the enzyme-specific pattern is stated in the certificate of analysis.
Absence of exonuclease activity
Approximately 5 μg [3H] labeled calf thymus DNA are incubated with 3 μl Not I for 4 hours at +37°C in a total volume of 100 μl 50 mM Tris-HCl, 10 mM MgCl2, 1 mM Dithioerythritol, pH approximately 7.5. Under these conditions, no release of radioactivity is detectable, as stated in the certificate of analysis.
Charakterystyka techniczna
Prokaryotic genomic DNA: Not I fragments are between 20 and 1,000 kb, depending on the GC content.
Yeast genomic DNA: Not I fragments are, on average, 200 kb.
Mammalian genomic DNA: Not I fragments are approximately 1,000 kb.
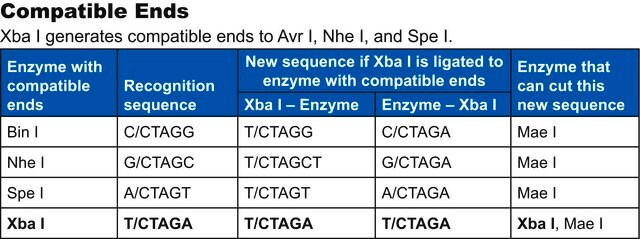
Compatible ends
Not I ends are compatible with ends generated by Eae I and EclX I (Xma III).
Isoschizomers
The enzyme has no known isoschizomers.
Methylation sensitivity
Not I is inhibited by the presence of 5-methylcytosine at the sites indicated (*) on the recognition sequence. However, the presence of 5-methylcytosine in the 5′-C position (°) is not inhibiting.
Relative activity in complete PCR mix
Relative activity in PCR mix (Taq DNA Polymerase buffer) is 0%. The PCR mix contained λDNA, primers, 10 mM Tris-HCl (pH 8.3, +20°C), 50 mM KCl, 1.5 mM MgCl2, 200 μM dNTPs, 2.5 U Taq DNA polymerase. The mix was subjected to 25 amplification cycles. After addition of 100 mM NaCl to the RE digest in the PCR mix, the activity of Not I still remains very low with below 5%.
Incubation temperature
+37°C
PFGE tested
Not I has been tested in Pulsed-Field Gel Electrophoresis (on bacterial chromosomes). For cleavage of genomic DNA (E. coli C 600) embedded in agarose for PFGE analysis, we recommend using 10 U of enzyme/μg DNA and 4 hour incubation time.
Ligation and recutting assay
Not I fragments obtained by complete digestion of 1 μg Ad2 DNA are ligated with 1 U T4 DNA Ligase in a volume of 10 μl by incubation for 16 hours at +4°C in 66 mM Tris-HCl, 5 mM MgCl2, 5 mM Dithiothreitol, 1 mM ATP, pH 7.5 (at +20°C) resulting in >80% recovery of Ad2 DNA.
Subsequent re-cutting with Not I yields >90% of the typical pattern of Ad2 × Not I fragments.
Profil DNA
- λ: 0
- φX174: 0
- Ad2: 7
- M13mp7: 0
- pBR322: 0
- pBR328: 0
- pUC18: 0
- SV40: 0
Definicja jednostki
Komentarz do analizy
The buffer in bold is recommended for optimal activity
- A: 10-25%
- B: 50-75%
- H: 100%
- L: 0-10%
- M: 25-50%
Inne uwagi
Tylko elementy zestawu
- Enzyme Solution
- SuRE/Cut Buffer H 10x concentrated
Kod klasy składowania
12 - Non Combustible Liquids
Klasa zagrożenia wodnego (WGK)
WGK 1
Temperatura zapłonu (°F)
does not flash
Temperatura zapłonu (°C)
does not flash
Certyfikaty analizy (CoA)
Poszukaj Certyfikaty analizy (CoA), wpisując numer partii/serii produktów. Numery serii i partii można znaleźć na etykiecie produktu po słowach „seria” lub „partia”.
Masz już ten produkt?
Dokumenty związane z niedawno zakupionymi produktami zostały zamieszczone w Bibliotece dokumentów.
Produkty
Termin "enzym restrykcyjny" wywodzi się z badań nad fagiem λ (fagiem lambda) Enterobacteria w laboratoriach Wernera Arbera i Matthew Meselsona.
The term “Restriction enzyme” originated from the studies of Enterobacteria phage λ (lambda phage) in the laboratories of Werner Arber and Matthew Meselson.
Nasz zespół naukowców ma doświadczenie we wszystkich obszarach badań, w tym w naukach przyrodniczych, materiałoznawstwie, syntezie chemicznej, chromatografii, analityce i wielu innych dziedzinach.
Skontaktuj się z zespołem ds. pomocy technicznej