T3411
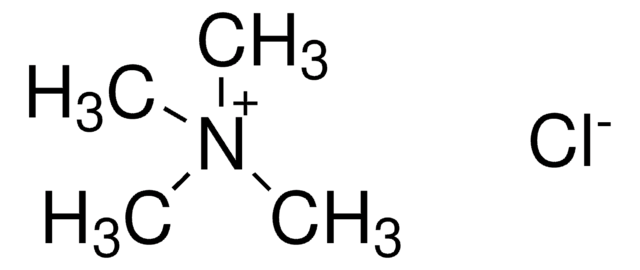
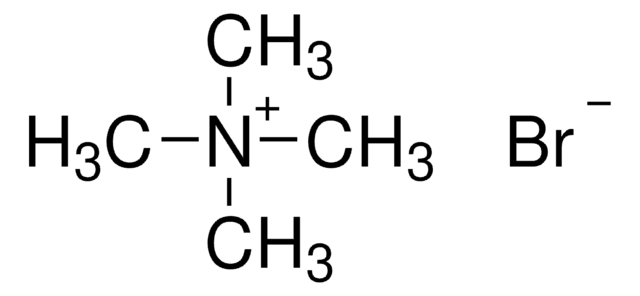
Tetramethylammonium chloride solution
for molecular biology
Sinonimo/i:
N,N,N-Trimethylmethanaminium chloride
Autenticatiper visualizzare i prezzi riservati alla tua organizzazione & contrattuali
About This Item
Prodotti consigliati
Grado
for molecular biology
Livello qualitativo
Concentrazione
5 M
Attività estranea
DNase, RNase, none detected
Stringa SMILE
[Cl-].C[N+](C)(C)C
InChI
1S/C4H12N.ClH/c1-5(2,3)4;/h1-4H3;1H/q+1;/p-1
OKIZCWYLBDKLSU-UHFFFAOYSA-M
Descrizione generale
Tetramethylammonium binds AT-rich DNA polymers while concomitantly abolishing the preferential melting of AT versus GC base pairs. It is supplied as a 0.2 μm filtered solution in 18 megohm water.
Applicazioni
Tetramethylammonium chloride solution (TMAC) has been used:
- in the preparation of hybridization cocktail for array hybridization and scanning
- in next-generation sequencing (NGS), and genome-wide unbiased identification of double-stranded breaks enabled by sequencing (GUIDE-seq) library preparation
- in the preparation of TMAC buffer and bead hybridization mixture for hybridization and detection
Certificati d'analisi (COA)
Cerca il Certificati d'analisi (COA) digitando il numero di lotto/batch corrispondente. I numeri di lotto o di batch sono stampati sull'etichetta dei prodotti dopo la parola ‘Lotto’ o ‘Batch’.
Possiedi già questo prodotto?
I documenti relativi ai prodotti acquistati recentemente sono disponibili nell’Archivio dei documenti.
I clienti hanno visto anche
W B Melchior et al.
Proceedings of the National Academy of Sciences of the United States of America, 70(2), 298-302 (1973-02-01)
Several small alkylammonium ions can eliminate, or even reverse, the usual dependence of the DNA transition temperature on base composition. For example, in 3 M tetramethylammonium chloride, or 2.4 M tetraethylammonium chloride, DNAs of different base compositions all melt at
Nikolay L Malinin et al.
Nature protocols, 16(12), 5592-5615 (2021-11-14)
Genome-wide unbiased identification of double-stranded breaks enabled by sequencing (GUIDE-seq) is a sensitive, unbiased, genome-wide method for defining the activity of genome-editing nucleases in living cells. GUIDE-seq is based on the principle of efficient integration of an end-protected double-stranded oligodeoxynucleotide
Hybridization of genomic DNA to oligonucleotide probes in the presence of tetramethylammonium chloride.
A G DiLella et al.
Methods in enzymology, 152, 447-451 (1987-01-01)
Xiao-Yong Li et al.
Methods in molecular biology (Clifton, N.J.), 809, 3-26 (2011-11-25)
Immunoprecipitation of cross-linked chromatin in combination with microarrays (ChIP-chip) or ultra high-throughput sequencing (ChIP-seq) is widely used to map genome-wide in vivo transcription factor binding. Both methods employ initial steps of in vivo cross-linking, chromatin isolation, DNA fragmentation, and immunoprecipitation.
Markus Welcker et al.
Genes & development, 27(23), 2531-2536 (2013-12-04)
The Fbw7 tumor suppressor targets a broad network of proteins for ubiquitylation. Here we show critical functions for Fbw7 dimerization in regulating the specificity and robustness of degradation. Dimerization enables Fbw7 to target substrates through concerted binding to two suboptimal
Il team dei nostri ricercatori vanta grande esperienza in tutte le aree della ricerca quali Life Science, scienza dei materiali, sintesi chimica, cromatografia, discipline analitiche, ecc..
Contatta l'Assistenza Tecnica.