おすすめの製品
詳細
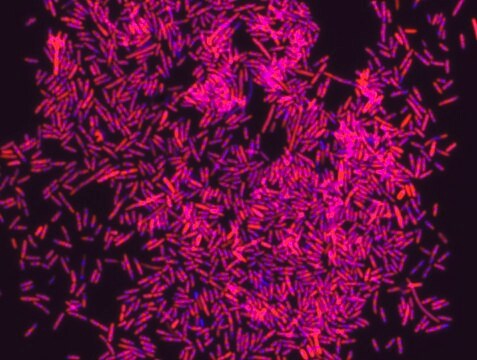
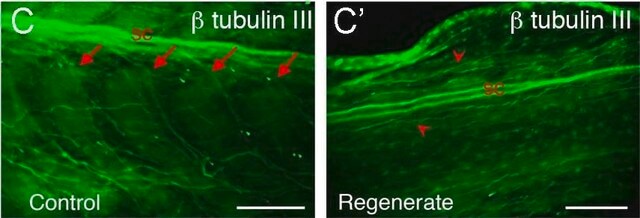
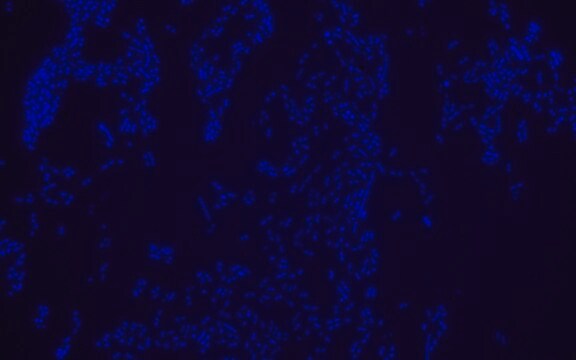
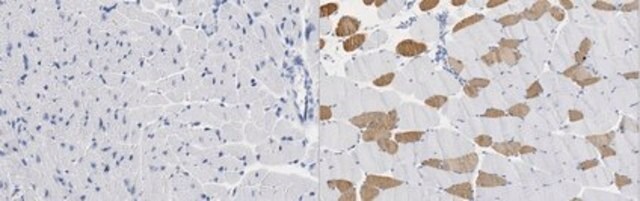
Fluorescent In Situ Hybridization technique (FISH) is based on the hybridization of fluorescent labeled oligonucleotide probe to a specific complementary DNA or RNA sequence in whole and intact cells. Microbial FISH allows the visualization, identification and isolation of bacteria due to recognition of ribosomal RNA also in unculturable samples. FISH technique can serve as a powerful tool in the microbiome research field by allowing the observation of native microbial populations in diverse microbiome environments, such as samples from human origin (blood and tissue), microbial ecology (solid biofilms, aquatic systems) and plants. It is strongly recommended to include positive and negative controls in FISH assays to ensure specific binding of the probe of interest and appropriate protocol conditions. We offer positive (MBD0032/33) and negative control (MBD0034/35) probes, that accompany the specific probe of interest. Enterococcus faecium probe specifically recognizes E. faecium cells,. E. faecium a gram-positive bacterium and is a cause of a variety of infections, including endocarditis, urinary tract infections, prostatitis, intra-abdominal infection, cellulitis, and wound infection as well as concurrent bacteremia. Studies have shown that E. faecium is highly resistant to multiple antibiotics and that vancomycin-resistant Enterococcus faecium (VRE) can be asymptomatically carried by healthy people, which can hamper hospitals’ attempts to control the spread of the bacteria. Over the past two decades, Enterococcus faecium has emerged as a leading cause of multidrug-resistant enterococcal infection in the United States. E. faecium has a high antibiotic-resistance with more than half of its pathogenic isolates expressing resistance to vancomycin, ampicillin, and high-levels of aminoglycosides.
アプリケーション
Probe for fluorescence in situ hybridization (FISH), recognizes Enterococcus faecium cells
特徴および利点
- Visualize, identify, and isolate E. faecium cells.
- Observe native E. faecium cell populations in diverse microbiome environments.
- Specific, sensitive, and robust identification of E. faecium in bacterial mixed population.
- Specific, sensitive, and robust identification even when E. faecium is in low abundance in the sample.
- FISH can complete PCR based detection methods by avoiding contaminant bacteria detection.
- Provides information on E. faecium morphology and allows to study biofilm architecture.
- Identify E. faecium in clinical samples such as tumor and brain tissues (for example in formalin-fixed paraffin-embedded (FFPE) samples), saliva and oral cavity and medical equipment such as dental implants and voice prostheses.
- The ability to detect E. faecium in its natural habitat is an essential tool for studying host-microbiome interaction.
保管分類コード
12 - Non Combustible Liquids
WGK
WGK 1
引火点(°F)
Not applicable
引火点(℃)
Not applicable
適用法令
試験研究用途を考慮した関連法令を主に挙げております。化学物質以外については、一部の情報のみ提供しています。 製品を安全かつ合法的に使用することは、使用者の義務です。最新情報により修正される場合があります。WEBの反映には時間を要することがあるため、適宜SDSをご参照ください。
Jan Code
MBD0047-50UL-PW:
MBD0047-50UL:
MBD0047-VAR:
最新バージョンのいずれかを選択してください:
ライフサイエンス、有機合成、材料科学、クロマトグラフィー、分析など、あらゆる分野の研究に経験のあるメンバーがおります。.
製品に関するお問い合わせはこちら(テクニカルサービス)