Enzyme Activity Assays
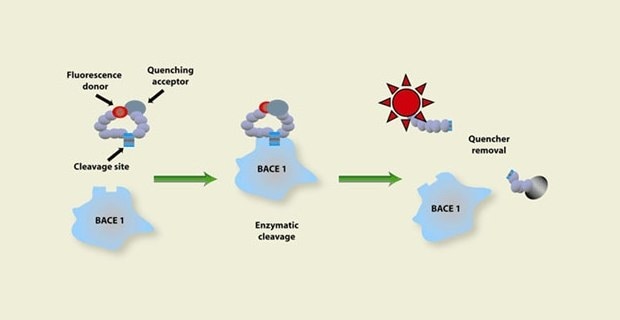
Figure 1. Representation of the enzymatic activity assay process starting with a fluorescence donor and quenching acceptor on a substrate going through enzymatic cleavage to remove the quencher, showing an increase in fluorescent activity which signals there is enzyme activity happening.
Enzymatic activity assays are predominately performed by researchers to identify the presence or quantity of a specific enzyme in an organism, tissue, or sample. Examples of such enzymes include α-amylase, catalase, laccase, peroxidase, lysozyme, and reporter enzymes alkaline phosphatase, and luciferase. Diverse reagents and methodologies are widely available that allow for the investigation of specific enzyme-substrate interactions. The selection of an appropriate workflow solution depends on the sensitivity that is required by the researcher. Colorimetric solutions are useful for detection, while fluorescence-based reagents are better suited for quantification of enzyme activity.
Featured Categories
Tissue dissociation and cell detachment resources, protocols, and a comprehensive offering of reliable and extensively characterized enzymes including, trypsin, collagenase, papain, nucleases (DNase and RNase), hyaluronidase, elastase, and protease XIV.
Substrates and enzymes are crucial in life science research, both as tools and targets in detection systems. Discover enzyme-based protein detection systems for ELISA, immunohistochemistry, western blotting, and many more with our diverse portfolio of detection substrates and enzymes.
Drug analysis enzymes, including β-Glucuronidase (beta-Glucuronidase) and sulfatase for drug metabolism and drug screening studies
Our portfolio of high-quality coenzymes reliably facilitate the function of enzymes and catalyze reactions for your research applications.
Determining Optimal Conditions for Enzymatic Activity
While numerous enzyme assays have been described in the literature, it is necessary to modify these procedures to the unique requirements of the enzyme under investigation. The specific activity of an enzyme depends on numerous factors including, pH, temperature, ionic strength, and the concentration of all components in the assay. For conditions such as pH, enzyme activity often resembles a bell curve. The highest activity at a specific pH is reported as the Vmax, with decreasing activity observed on either end of the curve. Some enzymes may require additional considerations for compounds that are not directly involved with the reaction, such as metal ions, detergents, and hydrophobic molecules.
Determining Optimal Conditions for Enzymatic Activity
While numerous enzyme assays have been described in the literature, it is necessary to modify these procedures to the unique requirements of the enzyme under investigation. The specific activity of an enzyme depends on numerous factors including, pH, temperature, ionic strength, and the concentration of all components in the assay. For conditions such as pH, enzyme activity often resembles a bell curve. The highest activity at a specific pH is reported as the Vmax, with decreasing activity observed on either end of the curve. Some enzymes may require additional considerations for compounds that are not directly involved with the reaction, such as metal ions, detergents, and hydrophobic molecules.
Enzyme Assay Components
For many enzymes, water is the standard solvent as it is the same water-based environment found in cells. However, organic solvents are used in some cases where enzymes or enzyme components are insoluble in water. Additionally, substrates and cofactors, catalysis for enzymatic reactions, are also critical components in an enzyme activity assay. Substrates and cofactors are often identified based on their functions under physiological conditions. As such, substrates and cofactors broadly interact and may be required by a diverse range of enzymes. Buffers and ions are additional critical elements to consider as they are essential for stabilizing the pH throughout the assay and directly influence enzyme activity. For example, mono- or divalent metal ions may be required for the catalytic activity of cofactors in the reaction and are essential for enzyme activity.
Performing an Enzyme Assay
The preparation of enzyme assay components is a practical first step. It is generally advantageous to prepare a large assay mixture, excluding one activating component, to avoid pipetting error that is inherent when pipetting small volumes. Once the assay mixture is prepared, researchers will then add the final activating component to the mixture to initiate the activity assay. Enzyme pretreatment by storing the enzyme at cool temperatures and often including various chemical or protein additives are critical factors to ensure enzyme stability and maximum activity before the initiation of the assay. Once the reaction materials are combined into an observation vessel, all the components should be quickly and thoroughly mixed as the reaction begins. Data recording should begin immediately after mixing, and the detectable signal from the assay should be plotted for the complete time course of the reaction.
Visit our document search for data sheets, certificates and technical documentation.
Related Articles
- Learn about compound library screening and explore our compound libraries, including our Library of Pharmacologically Active Compounds (LOPAC®1280) portfolio to experience the gold-standard collection of biologically annotated, ready-to-screen compounds.
- Cholesterol undergoes esterification to improve transport. Cholesterol esters are more easily packaged into the interior of lipoproteins - increasing the quantity that can be readily transported in the blood stream.
- Complete List of Enzyme Commission Numbers for metabolics research.
- This procedure applies to all products that have a specification for Cathepsin B activity determined by the liberation of 7-amino-4-methylcoumarin from Z-Arg-Arg 7-amido-4-methylcoumarin.
- Proteinase K (EC 3.4.21.64) activity can be measured spectrophotometrically using hemoglobin as the substrate. Proteinase K hydrolyzes hemoglobin denatured with urea, and liberates Folin-postive amino acids and peptides. One unit will hydrolyze hemoglobin to produce color equivalent to 1.0 μmol of tyrosine per minute at pH 7.5 at 37 °C (color by Folin & Ciocalteu's Phenol Reagent).
- See All (36)
Related Protocols
- This procedure applies to all products that have a specification for Cathepsin B activity determined by the liberation of 7-amino-4-methylcoumarin from Z-Arg-Arg 7-amido-4-methylcoumarin.
- This page provides information about different pull-down assays for the further isolation of multiprotein complexes to identify their components with products from Cytiva.
- Follow our procedure for the determination of alpha-Amylase activity. This enzymatic assay of a-Amylase guides you through the entire process and necessary calculations.
- This procedure may be used for all Catalase products.
- This procedure may be used for Hyaluronidase products.
- See All (95)
Find More Articles and Protocols
How Can We Help
In case of any questions, please submit a customer support request
or talk to our customer service team:
Email custserv@sial.com
or call +1 (800) 244-1173
Additional Support
- Chromatogram Search
Use the Chromatogram Search to identify unknown compounds in your sample.
- Calculators & Apps
Web Toolbox - science research tools and resources for analytical chemistry, life science, chemical synthesis and materials science.
- Customer Support Request
Customer support including help with orders, products, accounts, and website technical issues.
- FAQ
Explore our Frequently Asked Questions for answers to commonly asked questions about our products and services.
To continue reading please sign in or create an account.
Don't Have An Account?



