Viral RNA Extraction Buffer for SARS-CoV-2 in Saline and Saliva
- Extraction and Detection of SARS-CoV-2 with Viral RNA Extraction Buffer (VRE100)
- Viral RNA Extraction and qRT-PCR Amplification Method
- Viral RNA Extraction Buffer (VRE100) Sensitivity using qRT-PCR
- Viral Extraction Buffer (VRE100) Performance Comparison
- SARS-CoV-2 Viral RNA Extraction and LAMP Assay Analysis
- Benefits of Viral RNA Extraction Buffer for qRT-PCR and LAMP Assay Analysis
- RNA Detection of Influenza and RSV With VRE100
- VRE100 Viral RNA Extraction Buffer FAQ
- Product Ordering Information
The COVID-19 pandemic drastically elevated the need for rapid, reliable, and affordable viral detection techniques. Current approaches often depend on methods or reagents that have faced supply chain shortages, at times resulting in severe backlogs. Furthermore, silica-based RNA purification can be time-consuming, expensive, and challenging to automate. In addition to RNA isolation, sample collection via nasopharyngeal (NP) swab can be challenging, especially for children or cohorts that need to be routinely tested. Collection of saliva is an easier and less invasive process than NP swabs, and SARS-CoV-2 viral particles are stable in saliva for several days without any transport medium 1.
For Research Use Only. Not For Use In Diagnostic Procedures.
Extraction and Detection of SARS-CoV-2 and other respiratory RNA VIRUSES with Viral RNA Extraction Buffer (VRE100)
Here we demonstrate sensitive detection of crosslink-inactivated SARS-CoV-2 viral particles in both saline and human saliva through lysis with a novel Viral RNA Extraction Buffer (VRE100). The workflow consists of a 5-minute room temperature incubation which circumvents the need for RNA purification and cleanup. Samples are immediately ready for direct RNA analysis using molecular detection techniques, including qRT-PCR and Loop-Mediated Isothermal Amplification (LAMP) assays. Additionally, Viral RNA Extraction Buffer can detect other respiratory RNA viruses such as Influenza A, Influenza B, and Respiratory Syncytial Virus (RSV) in Viral Transport Medium.
VIRAL RNA EXTRACTION AND QRT-PCR AMPLIFICATION METHOD
Crosslink-inactivated viral particles (ZeptoMetrix® Corp.) were spiked into saline, saliva from healthy individuals, or transport medium at various concentrations. For lysis with Viral RNA Extraction Buffer (VRE100), saliva samples were centrifuged for 5 min at 15,000 RCF to remove particulates and supernatant was used for testing. 2 µl of VRE100 was added to 6 µl sample and incubated for 5 minutes at room temperature for SARS-Cov-2 or 15 minutes for Influenza A/B and RSV. For ProK + heat treatment, samples were spiked with Proteinase K to a final concentration of 2.4 mg/ml, vortexed for 1 min, heated at 95°C for 5 min, and briefly centrifuged to remove supernatant. 12 µl qRT-PCR Master Mix (QR0200) containing target-specific primers and probe was added directly to the RNA-extracted sample and analyzed on a BioRad CFX Connect™ instrument. qRT-PCR reaction conditions are shown below. For Research Use Only. Not For Use In Diagnostic Procedures.
Primers and Probes Used
SARS-CoV-2 Cycling Parameters:
Reverse Transcription 44 °C for 30m
Denaturation 94 °C for 2m
qPCR (45 cycles) 94 °C for 15s; 60 °C for 30s
Influenza A/B Cycling Parameters:
Reverse Transcription 44 °C for 30m
Denaturation 94 °C for 3m
qPCR (45 cycles) 94 °C for 15s; 55 °C for 15s; 72 °C for 30s
RSV Cycling Parameters:
Reverse Transcription 44°C for 30m
Denaturation 94 °C for 3m
qPCR (45 cycles) 94 °C for 15s; 47 °C for 15s; 72 °C for 30s
Viral RNA Extraction Buffer (VRE100) sensitivity using qRT-PCR
Saline is frequently available as a storage medium for NP swabs, and saliva offers a promising alternative to sample collection. Crosslink-inactivated SARS-CoV-2 particles were spiked into either saline or saliva from four different individuals at various concentrations based on manufacturer reported values. Samples were then treated with Viral RNA Extraction Buffer (VRE100) and detected by qRT-PCR (Figure 1). To test compatibility with several commonly used transport media, crosslink-inactivated SARS-CoV-2 particles were spiked at 1X104 particles per mL into indicated medium, samples were either left untreated or treated with Viral RNA Extraction Buffer, and RNA was detected by qRT-PCR (Figure 2). For Research Use Only. Not For Use In Diagnostic Procedures.
- Denaturation 94°C for 2m
- qPCR (45 cycles) 94°C for 15s; 60°C for 30s

Figure 1.Viral RNA Extraction Buffer (VRE100) sensitivity using qRT-PCR. A, Ct values obtained with decreasing amounts of viral particles in saline. B, Ct values obtained with decreasing amounts of viral particles spiked into human saliva. Each color represents saliva from a different healthy individual.
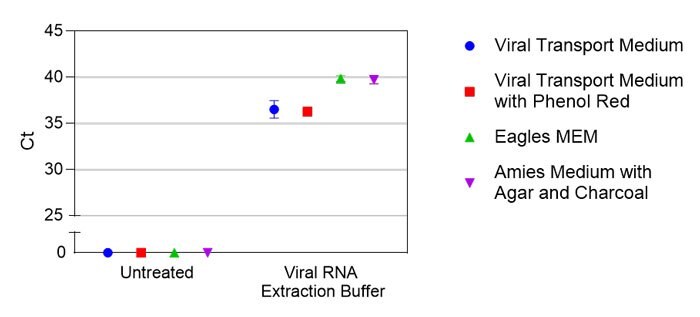
Figure 2.Comparison of virus detection in various transport media using Viral RNA Extraction Buffer (VRE100). Ct values are shown for treated or untreated samples in different media. Each condition was tested in duplicate.
Viral Extraction Buffer (VRE100) Performance Comparison
To test comparability to a similar approach, crosslink-inactivated SARS-CoV-2 particles were spiked into saliva from four different individuals at various concentrations. Each sample was subjected either to lysis with VRE100, treatment with Proteinase K and heat, or left untreated, and all samples were amplified by qRT-PCR (Figure 3).

Figure 3.Viral RNA Extraction Buffer performance relative to similar techniques. A, Ct values from four different saliva samples. Each color represents saliva from a different individual containing spiked viral particles. B, Amplification curves from a representative saliva sample (each treatment analyzed in duplicate).
SARS-CoV-2 Viral RNA Extraction and LAMP Assay Analysis
Colorimetric LAMP technology allows for a quick, simple assay readout that circumvents the need for qPCR instrumentation. Healthy human saliva was spiked with crosslink-inactivated SARS-CoV-2 (ZeptoMetrix® Corp.) at 5x104 viral particles per mL. 1.6 µl Viral RNA Extraction Buffer was added to 4.9 µl sample and allowed to incubate for 5 minutes at room temperature. Viral RNA was detected with a SARS-CoV-2 Rapid Colorimetric LAMP Assay Kit (NEB) as per manufacturer instructions, with heating for 40 minutes at 65°C, and results were determined by visual inspection (Figure 4). Synthetic SARS-CoV-2 RNA (1x105 copies, Twist Bioscience) was used as a positive assay control.

Figure 4.Viral RNA Extraction Buffer coupled with LAMP. Pink color indicated a negative result, while yellow indicated a positive result in which SARS-CoV-2 RNA was detected. Conditions with (+) or without (-) Viral RNA Extraction Buffer.
Benefits of Viral RNA Extraction Buffer for qRT-PCR and LAMP Assay Analysis
Viral RNA Extraction Buffer (VRE100) was effective in lysing inactivated SARS-CoV-2 particles at concentrations as low as 1x103 viral particles per mL in saline and 5x104 viral particles per mL in saliva. Additionally, 1.6x104 crosslink-inactivated Influenza A or RSV particles per mL, and 3x103 crosslink-inactivated Influenza B particles per mL, were detectable in Viral Transport Medium upon treatment with VRE100. Compatibility with various commonly used viral transport media was demonstrated at 1x104 viral particles per mL. The workflow was compatible with analysis via both qRT-PCR and LAMP. Detection of SARS-CoV-2 in saliva with Viral RNA Extraction Buffer was more consistent and sensitive than treatment with Proteinase K and heat. The use of Viral RNA Extraction Buffer also saved time and cost compared to comparable methods requiring RNA cleanup. These results demonstrate that Viral RNA Extraction buffer can provide a quick and convenient workflow for the detection of viral particles in saline, saliva, or viral transport media.
RNA Detection of Influenza and RSV with VRE100
Influenza and RSV continue to be significant public health concerns. Crosslink-inactivated Influenza A, Influenza B, or RSV particles were spiked into Viral Transport Medium at varying concentrations based on manufacturer reported values. Samples were then treated with VRE100 or left untreated, and subsequently detected by qRT-PCR (Figure 5). For Research Use Only. Not For Use In Diagnostic Procedures.
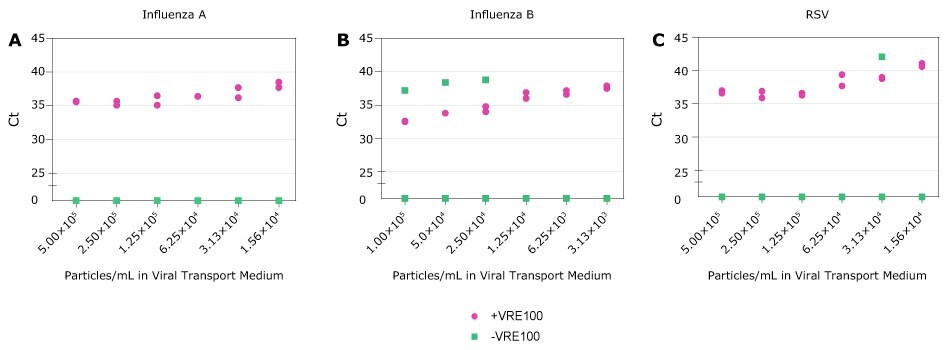
Figure 5 Detection of RNA Viruses. A-C, Ct values for Influenza A, Influenza B, and RSV obtained with decreasing amounts of viral particles in Viral Transport Medium in the presence (purple) or absence (green) of Viral RNA Extraction Buffer. Each condition was run in duplicate. Samples not detected are represented as zero on the Y axis.
VRE100 Viral RNA Extraction Buffer FAQ
What ratio of VRE100 to sample is recommended?
VRE100 was developed as a 4x lysis solution, such that 1 part buffer is added to 3 volume equivalents of sample. However, improved detection sensitivity may be achieved in certain samples by increasing the amount of VRE100 used for sample lysis.
Can samples lysed with VRE100 be stored for future use?
Yes, treated samples can be stored at -20 °C until ready for RNA detection. Multiple freeze-thaws are not recommended.
Which viruses can be detected using VRE100?
RNA from SARS-CoV-2, Influenza A and B, and Respiratory Syncytial virus (RSV) have been detected using VRE100 to lyse crosslink-inactivated viral particles in Viral Transport Medium.
Which viral transport media can be used with VRE100?
VRE100 has been shown to be compatible with saline, including phosphate-buffered saline (PBS) and Hank’s Buffered Saline Solution (HBSS), as well as with standard viral transport medium containing 2% FBS. Transport media containing guanidine are not recommended for use with VRE100 as guanidine will interfere with direct nucleic acid detection techniques, such as RT-qPCR. VRE100 is also compatible with undiluted saliva.
Is VRE100 capable of inactivating virus?
Viral inactivation after incubation with VRE100 has not been tested.
Can VRE100 be used for nasal swabs?
VRE100 has not been tested for use with nasal swabs, and is a Research Use Only product.
Is there any reaction stoppage step required?
No reaction stoppage step is required. Treated samples can be kept at room temperature for up to 30 min. If analyzing RNA within 1-2 hours samples can be kept on ice, otherwise samples should be frozen until ready for RNA detection.
References
To continue reading please sign in or create an account.
Don't Have An Account?