Cas9-Mediated Recombineering Enhances Metabolic Production of Kdo2-Lipid A by E. coli
Abstract
CRISPR-Cas systems are a new generation of RNA-guided microbial genome engineering tools that allow targeted genome modifications, such as insertions, deletions, and point mutations, to be introduced with greater ease than ever before. In this article, we present an application of our novel E. coli CRISPR/Cas-mediated Lambda-Red (λ-Red) homologous recombination (HR) vector system, which facilitates gene editing through the homology-directed repair (HDR) of double-stranded DNA breaks (DSBs) created by Cas9 endonuclease, using either ssDNA or dsDNA as an editing template. We use this tool to perform multi-gene editing in E. coli for production of a clinically relevant compound known as Kdo2-lipid A.
Introduction
Bacterial genome engineering is used for many important biotechnological and biomedical purposes; better tools for manipulating microbial genomes are in high demand. E. coli, one of the most commonly engineered bacteria, has an active homologous recombination (HR) system, so chromosomal integration can be achieved simply by donating DNAs with homology arms to recombine at a target site. While simple, this method is extremely inefficient, as endogenous systems exhibit very low HR activity.1 Recombineering is a popular method that utilizes phage-derived proteins, the Lambda-Red (λ-Red) system, to increase the efficiency of recombination in E. coli.2 Single-stranded DNA (ssDNA) donors integrate most efficiently and allow multiple loci to be targeted at once, but require high-throughput screening for selection.3 Selection markers can be integrated into double-stranded DNA (dsDNA) donors, but excision of these markers requires a second step and creates scar sequences4 that can trigger unwanted recombination events. Engineering a complex strain using these methods requires multiple time-consuming rounds of recombineering.2
To solve this, we have developed a novel, dual-vector Cas9-mediated recombineering system that allows researchers to create scarless insertions, deletions, and replacements with incredibly high efficiency in E. coli. Until recently, CRISPR/Cas9-based gene editing methods5 have been overlooked as a microbial gene editing tool because most bacteria lack endogenous DSB repair systems, rendering Cas9 nuclease cleavage toxic.6 However, when used to mediate recombineering, Cas9’s cytotoxicity is advantageous, as it eliminates the need for selection markers or high-throughput screening.3 In Cas9-mediated recombineering, phage proteins facilitate the HDR of Cas9-induced DSBs using either ssDNA or dsDNA donors as repair templates; therefore, Cas9 cleavage will kill any cells that fail to recombine with the donor template and those cells that are successfully repaired will be rescued (Figure 1C). This method avoids the creation of scar sites, can be used in multiplexing, and is less laborious than previous methods.7 In this article, we used our novel E. coli CRISPR λ-Red HR system to engineer an industrial E. coli strain with enhanced production of a valuable biomedical research compound known as Kdo2-lipid A.

Figure 1. Schematic of Dual-Vector E. coli CRISPR λ-Red HR System. (A) Vector Map of Cas9 Lambda Red Homologous Recombination Plasmid for E. coli. (B) Vector Map of CRISPR HR gRNA Plasmids for Bacteria. The 20-bp target region of the gRNA is customized for each gene target of interest. (C) Diagram of lacZ Positive Control Experiment. This dual-vector system utilizes a kill or rescue scheme to bypass the need for antibiotic or high-throughput screening. Targeting lacZ in wild-type E. coli allows simple blue-white screening to identify edited mutant strains. Wild-type cells expressing our Cas9 λ-red plasmid are transformed with 3 conditions. Transformation with a non-targeting, negative control gRNA yields a blue lawn, because Cas9 is not cleaving at the lacZ locus and cells remain wild-type; cells transformed with only a lacZ-targeting gRNA are cleaved but not repaired, resulting in death; cells transformed with a lacZ-targeting gRNA and a donor DNA template are successfully edited at the lacZ locus, knocking out function of this gene and yielding white colonies.
3-deoxy-D-manno-octulosonic acid (kdo2-lipid A) is an essential component of the outer wall of Gram-negative bacteria. Kdo2-lipid A constitutes the active endotoxic center of lipopolysaccharide (LPS), which triggers a potent host immune response through the complex of toll-like-receptor 4 (TLR4) and myeloid differentiation protein. While LPS molecules are usually large in size, have micro-heterogeneity, and are difficult to detect and quantify in vitro or in vivo, Kdo2-lipid A molecules are small, have micro-homogeneity, and can be easily quantified by electrospray ionization mass spectrometry (ESI/MS). For these reasons, Kdo2-lipid A is a better stimulator than LPS when studying innate immune systems. However, Kdo2-lipid A has not to date been synthesized chemically, and bacteria that directly synthesize and accumulate Kdo2-lipid A do not exist. Therefore, such a bacterial strain must be constructed to make the production of Kdo2-lipid A viable.
Results
Kdo2-lipid A is synthesized via a pathway involving nine enzymes; it does not naturally accumulate in the cell due to enzymes that catalyze reactions that consume Kdo2-lipid A (Figure 2). Therefore, knocking out the right combination of these genes in the degradation pathways should generate a mutant E. coli strain that accumulates Kdo2-lipid A and is suitable for large-scale production.

Figure 2. Kdo-Lipid A Degradation Pathways. Many enzymes are involved in the degradation of Kdo2-lipid A. Genes encoding these enzymes are all potential candidates for knockout to enhance the accumulation of Kdo2-lipid A in E. coli strain W3110. Figure adapted from Wang, et al (8).
We used our E. coli CRISPR λ-Red HR system to generate four mutant strains by knocking out four different novel combinations of these competing genes in E. coli W3110. For each target gene, we designed separate 20-bp gRNAs and approximately 1 kb dsDNA donor templates. gRNA sequences were designed to conform to the NGG proximal adjacent motif (PAM) requirement of Cas9. When multiple candidate gRNAs were identified, we chose those that were complementary to sequences earlier in the coding sequence (CDS). Donor DNA templates were created by amplifying 500-bp homology arms on each side of the CDS, then fusing these two amplicons directly together via fusion PCR. The resulting dsDNA product is homologous to the target site but omits the CDS; upon homology-directed repair, this CDS deletion is integrated into the target site (Figure 3).
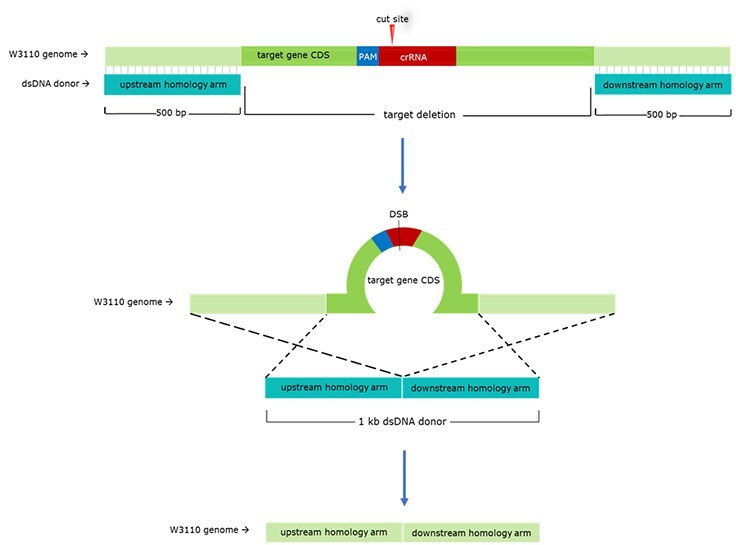
Figure 3. General Schematic for gRNA & dsDNA Donor Design. 500-bp homology arms are designed directly upstream and downstream of the target gene CDS. These two homology arms are fused via fusion PCR, creating a 1-kb dsDNA donor. Once Cas9 cuts at the gRNA-determined site, the lambda-red phage proteins will induce HDR (represented here by dotted lines) using the dsDNA donor as a repair template. Because the dsDNA donor omits the target gene CDS, HDR will result in a CDS-deleted mutant.
After the gRNA plasmids and donor templates were designed and cloned, they were transformed into an E. coli W3110 strain expressing our Cas9 λ-red HR plasmid. The Cas9 plasmid had been electroporated into E. coli W3110 in a previous step and was being maintained via antibiotic selection. These cells were then electroporated with gRNA plasmids and donor DNA templates corresponding to each target gene, setting up three reaction conditions following the same scheme shown in Figure 1C (Table 1, below). 24 hours after plating, candidate transformants were screened for deletion of target CDS via colony PCR (Figure 4) & Sanger sequencing (not shown). We used this protocol to generate two single knockout strains; we also used this protocol iteratively to generate two double-knockout strains.

Figure 4. Deletion in one of four novel W3110 E. coli strains engineered yields “fuzzy” colony phenotype characteristic of deletions at this locus (left). (A) Negative control plate of W3110 expressing pRedCas9 transformed with non-targeting guide RNA only yields ~107 CFU. (B) W3110 expressing pRedCas9 transformed with targeted gRNA without donor shows cytotoxicity, as expected, with ~103 CFU. (C) W3110 expressing pRedCas9 transformed with targeting gRNA and dsDNA donor with 500 bp homology arms yields ~105 CFU. (D) Colonies from plate C show “fuzzy” appearance. (E) Colony PCR confirms ~900 bp deletion in 7 out of 8 colonies compared to wild-type 2.1 kb PCR product. Genotype further confirmed via Sanger sequencing. Results (not shown) showed ≥95% of each target gene CDS was successfully deleted with no scarring of surrounding genome.
We then cultured each strain in 50-L fermentations to determine the viability of large-scale production of Kdo2-lipid A from each. After multiple purification steps, the purity of the resulting Kdo2-lipid A was ≥98%, as shown by the High-Performance Liquid Chromatography (HPLC) results in Figure 5A. We used Liquid Chromatography-Mass Spectrometry (LC-MS) to verify the identity of this product as Kdo2-Lipid A (Figure 5B). The post-purification yield from two of our novel strains is shown in Figure 6, below. Moving forward with only the strain corresponding to the highest yield, we then measured the activity of Kdo2-lipid A using a functional assay measuring TLR4 activation in Human Embryonic Kidney 293 (HEK) cells (Figure 7). Using the methods described here, we successfully identified a novel engineered strain suitable for large-scale production of Kdo2-lipid A.

Figure 5. Analysis of Product Isolated from Novel Engineered Strain. (A) HPLC analysis of Kdo2-lipid after purification. A. Novel strain produced high purity (≥98%) Kdo2-lipid A in a 50-L fermenter. The relative area – small peak is the impurity. Final product is ≥98% purity (B) LC-MS results verify Kdo2-lipid A exact mass. Mass spec analysis shows the mass observed is consistent with the expected mass of Kdo2-lipid A.

Figure 6. Kdo2-lipid A yield from two novel engineered strains in 50-L fermentation. Strain 2 generated a 14X yield increase over strain 1, making it the best candidate for large-scale production of Kdo2-lipid A. Note that there is no reliable baseline to calculate overall yield improvement compared to wild-type E. coli, because Kdo2-lipid A does not naturally accumulate in wild-type strains.
Discussion & Conclusion
In this simple gene-editing experiment, we successfully generated and identified an industrially viable Kdo2-lipid A production strain using our novel E. coli CRISPR λ-Red HR system. Using 1 kb dsDNA donors, we created four different single- and double-knockout strains, removing ≥95% of the CDS of each gene of interest in one simple step per gene. Because of the toxicity of unrepaired Cas9-induced DSBs, we easily identified repaired, donor-integrated colonies without the use of antibiotic cassettes or high-throughput screening. To identify the most industrially viable strain among our four candidates, we scaled each up to 50-L fermentations, then evaluated the resulting Kdo2-lipid A yield and activity. Kdo2-lipid A does not naturally accumulate in native strains, as it is quickly consumed by downstream reactions. By knocking out genes corresponding to these catalytic reactions, we generated and identified a novel strain with 14–70X increased yield of high-purity, high-activity Kdo2-lipid A, ultimately making Kdo2-lipid A an economically and industrially feasible product for commercialization.
References
To continue reading please sign in or create an account.
Don't Have An Account?