Proline Derivatives and Analogs
Introduction
Proline is a non-polar proteinogenic amino acid that forms a tertiary amide when incorporated into peptides. It does not have a hydrogen on the amide group and therefore cannot act as a hydrogen bond donor. Proline is known as a classical breaker of both the α-helical and β-sheet structures in proteins and peptides. Nevertheless, it is widely distributed in the putative transmembrane domains of many protein transporters and channels, regions believed to be α-helical.1
Among the proteinogenic amino acids, proline plays a special role. In protein structures the planar peptide bond occurs predominantly in the trans conformation.2 The proline residue restricts the conformational space of the peptide chain. However, due to the small free enthalpy difference between the cis and trans Xaa-Pro bond isomers of 2.0 kJ·mol–1 (compared to 10.0 kJ·mol-1 for other Xaa-non-Pro peptide bonds), there is a relatively high intrinsic probability of 30% cis conformation at RT and both cis and trans isomers are present in solution.3,4
The cis/trans-isomerization of peptide bonds on the N-terminal side of Pro residues plays a key role in the folding process of a protein because the rotational barrier of the cis/trans-isomerization is quite high (85,0 ± 10,0 kJ·mol–1). Therefore, this interconversion is described to be one of the limiting steps of protein folding in vitro and in vivo.5 In nature there is a class of enzymes, the peptidyl-prolyl-cis/trans-isomerases (PPIases). They are able to catalyze protein folding by accelerating the isomerization of the Xaa-Pro-bond.6–8
Comparative studies performed with proline analogues revealed that the key step in the catalysis of the cis/trans-isomerization of a peptidyl-prolyl bond is a reduction of the double bond character of the planar, conjugated C–N amide bond. Any factor that can weaken the double bond character of the amide bond by destabilizing the planar peptide bond, or shifting the hybridization of the prolyl nitrogen from sp2 to sp3, is expected to accelerate the isomerization.9,10
In order to understand the relationship between imide bond geometry and bioactivity of peptides,11,12 synthetic proline analogues have been developed that provide restrictions of the Xaa-Pro imide conformation. Such proline mimetics are based on ring substitutions with alkyl and aromatic groups, incorporation of heteroatoms into the ring, or the expansion or contraction of the proline ring (Table 1). Those analogues are promising candidates for conformational studies and for tuning the biological, pharmaceutical, or physicochemical properties of naturally occuring, as well as de novo designed, linear, and cyclic peptides.
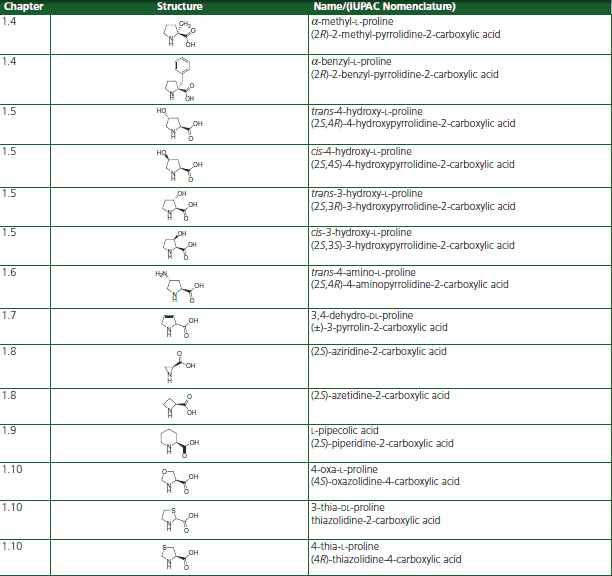
Table 1.Proline Analog or Homolog Structures for the Restriction of the Xaa-Pro Imide Conformation
Several proline analogs and homologs occur in nature. Trans-3-hydroxyproline and trans-4-hydroxyproline represent constituents of common proteins as a result of post-translational hydroxylation, especially in collagens.13 Various 3- and 4-alkylated derivatives of proline and hydroxyproline as well as analogues with ring restrictions, such as aziridine-2-carboxylic acid and azetidine-2-carboxylic acid, and ring expansions, i.e. pipecolic acid, are found in natural products.14,15 Derivatives such as L-azetidine-2-carboxylic acid, cis-4-hydroxy-L-proline, and 3,4- dehydro-DL-proline prevent pro-collagen from folding into a stable triple-helical conformation, thereby reducing excessive deposition of collagen in fibrotic processes and the growth of tumors.16
Thiazolidine-4-carboxylic acid thiaproline has also been incorporated into collagen model compounds17,18 and other bioactive molecules such as thrombin inhibitors,19 somatostatin,20,21 dipeptidyl peptidase IV substrates,22 angiotensin II,23 HIV inhibitors,24 ACE inhibitors,25 and oxytocin.26
α-Methyl-proline is a bioactive molecule restoring normal levels of bone collagen type I synthesis.27 It can be looked at as a conformationally constrained aminoisobutyric acid analog. The α-Methyl-proline residue has been inserted into morphiceptin to perform conformational studies on the bioactivity of the Xaa-Pro cis-/trans-isomers.28 A α-methyl-proline containing potential dual α4β1 integrin antagonist has been described recently.29
α-Benzyl-proline combines the conformational restrictions of a proline derivative with the electronic properties of phenylalanine. Spirolactams containing an α-benzyl-proline substructure have been synthesized as potential beta-turn mimetics.30
References
To continue reading please sign in or create an account.
Don't Have An Account?