Bioluminescent Firefly Luciferase Assays
What is Firefly Luciferase?
Firefly luciferase is a widely used bioluminescent reporter for studying gene regulation and function. It is a very sensitive genetic reporter due to the absence of endogenous luciferase activity in mammalian cells or tissues. Firefly luciferase is a 62,000 Dalton protein, which is active as a monomer and does not require subsequent processing for its activity. The enzyme catalyzes ATP-dependent D-luciferin oxidation to oxyluciferin, producing light emission centered at 560 nm. Light emitted from the reaction is directly proportional to the number of luciferase enzyme molecules.
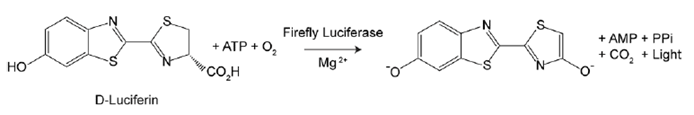
Figure 1. Luciferase assay principle. Bioluminescent reaction catalyzed by firefly luciferase produces light.
What is the Difference Between Bioluminescence and Fluorescence?
Bioluminescence is a chemical process in which an enzyme breaks down a substrate (such as Luciferase and D-luciferin) and one of the by-products of this reaction is light. Bioluminescence naturally occurs in nature in various algae, bacteria, fungi and some aquatic animals such as jellyfish. Fluorescence is a physical process by which light excites electrons in the fluorophore to a higher energy state, and when that electron falls back down to its ground state it emits a photon.
How do Luciferase Assays Work?
Firefly luciferase assays (SCT151, SCT154, LUC1, LUCASSY-RO) are designed for simple and efficient quantitation of firefly luciferase reporter enzyme activity from cultured cells with high sensitivity and linearity. This is a flash-type luminescence assay that requires signal to be measured immediately after adding working solution to samples. The luminescence signal decays over the course of about 10 minutes of reaction time, although signal half-life may vary depending on luciferase expression levels.
The light production resulting from the luciferase reaction leads to formation of suicidal adenyl-oxyluciferin at the enzyme surface. It results in very short half-life of the light emission with a flash-type kinetics. The Firefly Luciferase HTS Assay (SCT150) is designed with a proprietary mixture of substances that modify the enzymatic reaction to produce a long-lasting signal (steady-glow) by preventing the formation of adenyl-oxyluciferin at the enzyme surface. It is a homogeneous high sensitivity firefly luciferase reporter gene assay kit for the quantification of firefly luciferase expression in mammalian cells with signal half-life of about 3 hours (Figure 2). Glow-type luciferase assays have lower luminescence signal compared to flash-type assays. The sensitivity and limit of detection of the assay will depend on luciferase expression levels in your experimental system as well as luminometer sensitivity.

Figure 2.Titration of recombinant firefly luciferase in the firefly luciferase assay. A) Recombinant luciferase was serially diluted in 1X Firefly Lysis Buffer with 1 Mg/mL BSA and measured in the assay. B) The Firefly Luciferase HTS Assay is a steady-glow high sensitivity firefly luciferase reporter gene assay kit for the quantification of firefly luciferase expression in mammalian cells with signal half-life of about 3 hours.
What is the Difference Between Firefly and Renilla Luciferase?
Firefly luciferase assays uses luciferin in the presence of oxygen, ATP and magnesium to produce light (Green/Yellow, 550-70 nM), while Renilla luciferase assays (SCT153) requires only coelenterazine and oxygen to produce light (Blue, 480 nM). Renilla luciferase has been used as a reporter gene for studying gene regulation and function in vitro and in vivo. It commonly is used in multiplex transcriptional reporter assays or as a normalizing transfection control for firefly luciferase assays. The enzyme does not require post-translational modification for its activity and may function as a genetic reporter immediately following translation. Coelenterazine, substrate for Renilla luciferase, also emits light from enzyme-independent oxidation, a process known as autoluminescence. The autoluminescence is enhanced by superoxide anion and peroxynitrite in cells and tissues.
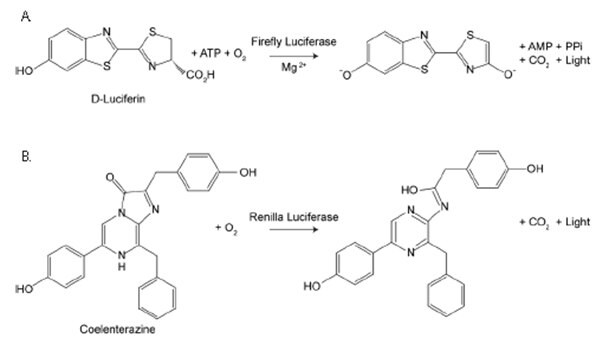
Figure 3.Firefly vs. Renilla Luciferase. Bioluminescent reactions catalyzed by firefly luciferase and Renilla luciferase.
What are the Advantages of Dual Luciferase Assays?
The Firefly/Renilla Dual Luciferase Assay (SCT152) allows measurement of both Firefly and Renilla luciferase activity in the same sample with high sensitivity and linearity. Firefly luciferase activity is measured first, then Renilla Luciferase Assay Buffer 2.0 is added to simultaneously quench firefly luciferase activity and measure Renilla luciferase activity. Renilla Luciferase Assay Buffer 2.0 quenches the firefly luciferase activity to the level of un-transfected cells, allowing sequential measurement of firefly and Renilla luciferase activity in the same sample. This is a flash-type assay that requires luminescence to be measured immediately after adding the detection reagents to the luciferase sample. Firefly signal decays over the course of about 12 minutes, while Renilla signal decays over the course of about 2 minutes, although this may vary depending on enzyme levels.


Figure 4.Dual luciferase assay overview. Example of Firefly & Renilla Luciferase detection using lysates from untransfected HeLa cells or cells transfected with either firefly luciferase alone (Firefly Only) or co-transfected with firefly and Renilla luciferases (Firefly + Renilla). In cells transfected with firefly only, the Renilla signal is the residual firefly luminescence after adding Renilla working solution to the reaction.
What are the Applications of Luciferase Assays?
Gene Reporter
Genetic reporters are used as indicators to study cellular events coupled to gene expression. Typically, a reporter gene is cloned with a DNA sequence of interest into an expression vector (which carries a Luciferase gene) that is then transferred into cells. Following transfer, the cells are assayed for the presence of the reporter by directly measuring the reporter protein itself or the enzymatic luciferase activity. A good reporter gene can be identified easily and measured quantitatively when it is expressed (in the organism or cells of interest).
For example, Sigma and SwitchGear Genomics™ have teamed up to exclusively offer a genome-wide collection of over 10,000 3′UTR regions in our optimized lentivirus luciferase reporter vector system. When conducting experiments using MISSION®3′UTR Lenti GoClones, including the proper controls permits accurate interpretation of the experimental gene expression results.

Cell Viability via ATP Detection
Because ATP is an indicator of metabolically active cells, the number of viable cells can be assessed based on the amount of ATP available. The ATP Cell Viability Luciferase Assay (SCT149, 11699709001) offers a highly sensitive homogenous assay for quantifying ATP in cell cultures. This kit takes advantage of Firefly luciferase’s use of ATP to oxidize D-Luciferin and the resulting production of light in to assess the amount of ATP available in cell cultures. The sensitive assay procedure involves a single addition of ATP detection cocktail directly to cells cultured in a serum-supplemented medium. No cell washing, medium removal and multiple pipetting are required. The kit can be used to detect as little as a single cell or 0.01 picomoles of ATP. The signal produced is linear within 6 orders of magnitude. By relating the amount of ATP to the number of viable cells, the assay has wide applications, ranging from the determination of viable cell numbers to cell proliferation to cell cytotoxicity.

Figure 5.ATP cell viability assay. Firefly luciferase’s use of ATP to oxidize D-Luciferin and the resulting production of light in order to assess the amount of ATP available that correlates to cell number and viability.
In vivo Imaging
Bioluminescence imaging (BLI) has become a popular technique for optical tracking of cells in small laboratory animals such as mice. Along with the low background autofluorescence and cellular toxicities, BLI can enable simultaneous visualization of monitoring for the expression of two divergent luciferase proteins by use of their specific substrates. The imaging dual luciferase gene (red codon optimized firefly luciferase and a green click beetle luciferase) activities in the same animals could reduce variations from individual differences of the experimental animals.

To continue reading please sign in or create an account.
Don't Have An Account?