11062590001
Roche
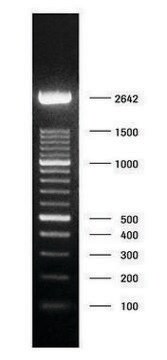
DNA Molecular Weight Marker VI
pkg of 50 μg (in 200 μl), solution
Synonym(s):
DNA marker
Sign Into View Organizational & Contract Pricing
All Photos(1)
About This Item
UNSPSC Code:
41105335
Recommended Products
form
solution
Quality Level
packaging
pkg of 50 μg (in 200 μl)
manufacturer/tradename
Roche
concentration
250 μg/mL
color
colorless
solubility
water: miscible
storage temp.
−20°C
General description
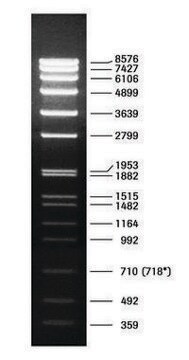
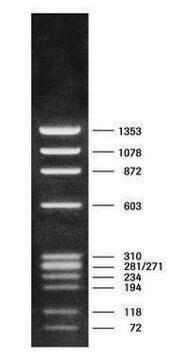
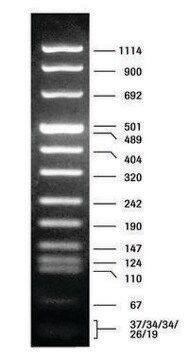
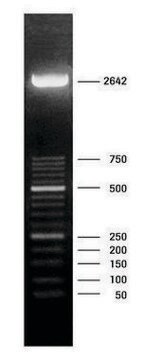
DNA Molecular Weight Marker VI is a fragment mixture prepared by cleavage of pBR328 DNA with restriction endonucleases Bgl I and pBR328 DNA digested with Hinf I. It has a size range of 0.15 to 2.1kbp.
The product is also available as DNA Molecular Weight Marker VI, Digoxigenin-labeled.
The product is also available as DNA Molecular Weight Marker VI, Digoxigenin-labeled.
Specificity
Specificity: Product has 5′-phosphorylated ends (restriction fragments).
Restriction site: Claevage sites and corresponding fragment lengths of pBR328:
- Bgl I cut sites at: 935, 1169, 2399, 4575, 4809 => creating following fragment sizes, respectively: 234 bp, 1230 bp, 2176 bp, 234 bp, 1033 bp.
- Hinf I cut sites at: 632, 852, 1006, 1304, 1757, 2274, 4040, 4434, 4732, 4886. =>
creating following fragment sizes, respectively: 220 bp, 154 bp, 298 bp, 453 bp, 517 bp,
1766 bp, 394 bp, 298 bp, 154 bp, 653 bp
Restriction site: Claevage sites and corresponding fragment lengths of pBR328:
- Bgl I cut sites at: 935, 1169, 2399, 4575, 4809 => creating following fragment sizes, respectively: 234 bp, 1230 bp, 2176 bp, 234 bp, 1033 bp.
- Hinf I cut sites at: 632, 852, 1006, 1304, 1757, 2274, 4040, 4434, 4732, 4886. =>
creating following fragment sizes, respectively: 220 bp, 154 bp, 298 bp, 453 bp, 517 bp,
1766 bp, 394 bp, 298 bp, 154 bp, 653 bp
Application
DNA Molecular Weight Marker VI is used for the size determination of DNA in agarose gels. It simplifies accurate molecular weight determination of double-stranded DNA fragments generated by:
- Restriction digests
- PCR
- RT-PCR
Sequence
As determined by computer analysis of the pBR328 sequence, the mixture contains 15 DNA fragments with the following base pair lengths (1 base pair = 660 daltons) occuring once: 220, 394, 453, 517, 653, 1033, 1230, 1766, 2176 and fragments 154,234, and 298 occuring twice as a result of the digestion of pBR328 with both Bgl I and Hinf I.
Note: Fragment lengths are derived from computer analysis of the DNA sequence. Depending on the size range of the marker, the smallest fragments will only be visible on overloaded gels.
Note: Fragment lengths are derived from computer analysis of the DNA sequence. Depending on the size range of the marker, the smallest fragments will only be visible on overloaded gels.
Unit Definition
50 μg = 1A260 unit
Physical form
Ready-to-use solution, 250μg/ml, in TE buffer (10mM Tris-HCl, 1mM EDTA, pH 8.0).
Preparation Note
Storage conditions (working solution): 2 to 8 °C
Once thawed we recommend further storage at 2 to 8 °C.
Once thawed we recommend further storage at 2 to 8 °C.
Other Notes
For life science research only. Not for use in diagnostic procedures.
Storage Class
12 - Non Combustible Liquids
wgk_germany
nwg
flash_point_f
does not flash
flash_point_c
does not flash
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
G Meulemans et al.
Avian pathology : journal of the W.V.P.A, 30(6), 655-660 (2001-12-01)
A polymerase chain reaction combined with restriction enzyme analysis was developed for detection and differentiation of all 12 fowl adenovirus (FAdV) serotypes representing the five fowl adenovirus (A to E) species. For primer design, the published sequences of the hexon
Qualitative PCR?ELISA protocol for the detection and
typing of viral genomes
typing of viral genomes
Monica Musiani
Nature Protocols (2007)
R Ferretti et al.
Applied and environmental microbiology, 67(2), 977-978 (2001-02-07)
A PCR-based method for the detection of Salmonella spp. in food was developed. The method, set up on typical salami from the Italian region of Marche, is sensitive and specific and shows excellent correlation with the conventional method of reference
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service