UDGHL-RO
Roche
Uracil-DNA Glycosylase, heat-labile
recombinant from marine bacterium BMTU 3346
Synonym(s):
PCR, UDG
About This Item
Recommended Products
biological source
bacterial (marine bacterium BMTU 3346)
Quality Level
recombinant
expressed in E. coli
form
solution
packaging
pkg of 100 U (11775367001)
pkg of 500 U (11775375001)
manufacturer/tradename
Roche
concentration
1000 U/mL
technique(s)
activity assay: suitable
color
colorless
optimum pH
8.3-8.9
solubility
water: soluble
suitability
suitable for enzyme test
application(s)
life science and biopharma
foreign activity
DNA decontamination:, present ( >= 10 e5 copies )
RNases with MS- II- RNA ≤10 units, none detected
Unspec.nucleases w. lambda-DNA 20 units, none detected
Unspec.nucleases w.M13mp9ss-DNA ≤20 units, none detected
Unspec.nucleases w.pBR 322-DNA ≤20 units, none detected
storage temp.
−20°C
Related Categories
General description
Enzyme Characteristics
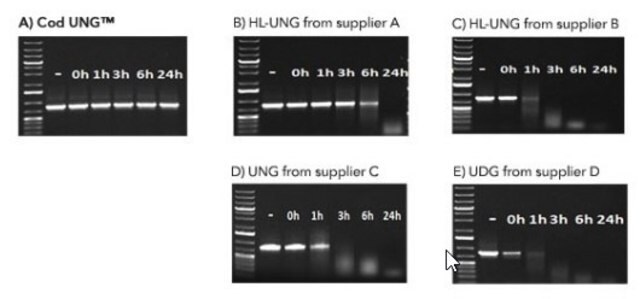
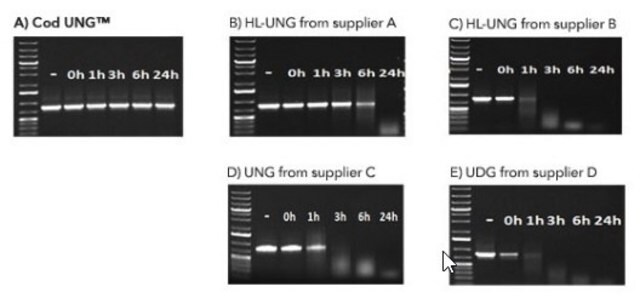
The BMTU 3346 enzyme is inactivated more quickly (2 minutes at +95°C) than the corresponding enzyme from E. coli (10 minutes at +95°C). It has also been reported that UNG from E. coli remains partially active, leading to the degradation of the dU-containing PCR product. In contrast, the heat-labile UNG does not degrade dU-PCR products within at least several hours of incubation at +2 to +8°C. Therefore, it is not necessary to freeze the PCR product immediately after amplification or to hold the reaction mixture at -70°C.
Specificity
- Uracil-DNA glycosylase hydrolyzes uracil-glycosidic bonds at U-DNA sites in single- and doublestranded DNA, excising uracil and creating alkali sensitive abasic sites in the DNA.
- The enzyme is active on both single-stranded DNA and double-standed DNA.
- Uracil-DNA glycosylase is inactive on RNA and native, uracil-free DNA.
- Since uracil-DNA glycosylase has no metal ion requirements, it is fully active in the presence of EDTA.
Heat inactivation: 95 °C for 2 min
Application
Important Note: For highly sensitive techniques like real-time PCR we recommend our LightCycler® Uracil-DNA Glycosylase which is optimized for this application.
Features and Benefits
- Prevent carryover contamination in PCR.
- Increase the efficiency of site-directed mutagenesis procedures.
- Label oligonucleotide probes.
- Perform faster inactivation than the corresponding enzyme from E. coli due to the lower thermostability of the enzyme.
- Obtain no degradation of dU-PCR products within at least several hours of incubation at +2 to +8°C.
Contents
The enzyme is supplied as 1 U/μl solution in storage buffer.
Quality
The enzyme does not contain any contaminating exo- or endonucleases and is free from RNase activity, according to the current quality control procedures.
Unit Definition
One Lindahl unit is defined as the amount of enzyme necessary to release 1 mol uracil at +37 °C in 1 minute. One Lindahl unit is comparable to 520 000 units based on our unit definition.
Volume Activity: 1 U/μl
Other Notes
Legal Information
Hazard Statements
Precautionary Statements
Hazard Classifications
Aquatic Chronic 3
Storage Class Code
12 - Non Combustible Liquids
WGK
WGK 2
Flash Point(F)
does not flash
Flash Point(C)
does not flash
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
Articles
DNA damage and repair mechanism is vital for maintaining DNA integrity. Damage to cellular DNA is involved in mutagenesis, the development of cancer among others.
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service