MSI Testing and IHC
Mike Lacey, M.D.
Medical Director
MSI Testing
History
Mismatch Repair
Prognostic Significance of Mismatch Repair
Deficiency in Colorectal Cancer
Genetic Counseling and Management of
Hereditary Nonpolyposis Colon Cancer
Prognosis
MSI Testing
Colorectal cancer is a common occurrence among inhabitants of most Western countries, second only to carcinoma of the lung.1 Despite improvements in surgical techniques and chemotherapeutic regimens, the prognosis of this disease has not improved significantly over the last decades. These concerns mandate greater attention to the etiology and pathogenicity of colorectal cancer.
Although dietary factors have been well researched in etiologic studies2, the most predictable risk factor of the disease, its hereditary predisposition, has received limited attention. This may be due to the paucity of knowledge that clinicians have about the role of genetic factors in the etiology of cancer.3 For example, many physicians believe that familial adenomatous polyposis (FAP) is the only genetic risk factor implicated in colorectal cancer’s etiology. However, FAP accounts for approximately 1% of the total colorectal cancer burden, whereas hereditary colorectal cancer syndromes are at least six to eight times as common as FAP.3,4
Hereditary nonpolyposis colorectal carcinoma (HNPCC), or Lynch syndrome, is an autosomal dominant syndrome accounting for 5 to 10% of the total colorectal cancer burden.5 The lack of characteristic diagnostic features has prompted the introduction of certain criteria to establish this diagnosis: (a) histologically verified colorectal cancer in at least three relatives (one of whom is a first-degree relative of the other two), (b) the presence of disease in at least two successive generations, and (c) one or more colorectal cancer cases diagnosed before the age of 50 years. These criteria, known as the “Amsterdam criteria I,” were proposed to identify HNPCC kindreds having hereditary colon carcinoma.5-7
Typical features of HNPCC include a family history of colorectal cancer (CRC) at a relatively young age, a predominance of proximal tumors, and a tendency to have multiple primary tumors, synchronous or metachronous. Certain types of extracolonic tumors are also associated with the disease, such as tumors in the endometrium, ovary, stomach, small bowel, hepatobiliary tract, pancreas, brain, and urinary tract.6,8-10
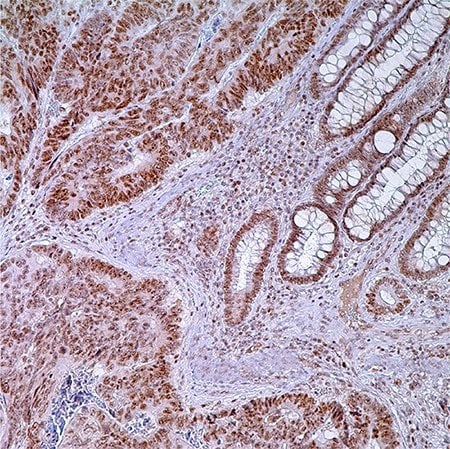
Figure 1. PMS2 (EPR3947†) - adenocarcinoma of colon. Note nuclear labeling of adenocarcinoma cells, normal crypt cells, and lymphocytes in lamina propria.
History
Henry T. Lynch came upon the disease that would ultimately bear his name through consultation on a patient with a strong history of CRC in the absence of overt polyposis.29,30 Pedigree analysis of that patient’s family formed the basis of Lynch’s 1966 “Hereditary factors in cancer. Study of 2 large midwestern kindreds’’, published in the Arch Intern Med.17 Lynch and Krush’s updating of the family, published in 1971, contains data on over 650 family members. Many of the syndrome’s salient features were noted: (1) increased incidence of adenocarcinomas, mainly of the colon and endometrium; (2) increased risk for multiple tumors; (3) autosomal dominant inheritance; and (4) early onset of cancer.10 Despite Lynch and Krush’s insight, their views met with considerable skepticism from the scientific community at large, as contemporary thinking was dominated by the notion that environmental factors were the principal determinant of cancer.29,30

Figure 2. Recommendations for genetic screening for hereditary nonpolyposis colorectal cancer (HNPCC).
Lynch’s ideas slowly gained traction, particularly in the international community, culminating in the organization of the “International Collaborative Group on Hereditary Non-Polyposis Colorectal Cancer’’ composed of “30 leading experts from 8 different countries’’. The group met in Amsterdam in the summer of 1990. The major output from that meeting was the formulation of a series of clinical criteria (now known as the Amsterdam criteria and colloquially known as the “3-2-1’’ rule) to serve as a common starting point for future research: (1) at least 3 relatives with histologically confirmed colorectal cancer, 1 of whom is a first degree relative of the other 2; familial adenomatous polyposis should be excluded; (2) at least 2 successive generations involved; and (3) at least 1 of the cancers diagnosed before age 50.19 The Amsterdam criteria (AC) were updated at the 1998 group meeting, acknowledging the significance of a subset of extracolonic tumors (ie, endometrium, small bowel, ureter, & renal pelvis). These revised AC-II are presented in Table 1.
Amsterdam Criteria II: Clinical Criteria for Families With HNPCC* (Table 1)
- ≥3 relatives with an HNPCC-associated cancer**
- ≥2 successive generations affected
- ≥1 diagnosed before the age of 50 years old
- 1 should be a first-degree relative of the other 2
- Familial adenomatous polyposis should be excluded
- Tumors should be verified by pathologic examination
Gastroenterology. 1999;116:1453–1456.2
*All criteria must be fulfilled.
**Colorectal cancer, cancer of the endometrium, small bowel, ureter, or renal pelvis. HNPCC indicates hereditary nonpolyposis colorectal cancer.
The molecular genetic basis of Lynch syndrome was largely elucidated in a rapid succession of reports from 1993 to 1994. Peltomaki and colleagues linked the disease (in 2 large kindreds meeting the original AC) to a locus on chromosome 2p. Then, 3 groups independently reported a peculiar molecular phenotype in CRC characterized by widespread alterations in the sequence length of simple repetitive sequences, a phenomenon, which the groups variously termed “replicative errors (RERs)’’, “MSI’’, and “ubiquitous somatic mutations in simple repetitive sequences’’. Aaltonen et al.31 found the RER-positive phenotype in 11/14 (79%) “hereditary nonpolyposis colon cancer (HNPCC) tumors’’ and 6/46 (13%) sporadic CRC. The sporadic RER+ tumors shared a right-sided predominance and near diploid status with the HNPCC cases. Thibodeau et al.32 found some degree of MSI in 25/90 (28%) CRC, linking this phenomenon to proximal location and to improved survival and demonstrating an inverse relationship with loss of heterozygosity. Ionov’s group reported ubiquitous somatic mutations in simple repetitive sequences in 12% of CRC, which was relatively more common in women and was associated with right-sided location, poorly differentiated histology, fewer KRAS and p53 mutations, and lower tumor stage.142 By the end of 1993 the relevant gene on 2p, MSH2, had been cloned and germline mutations identified in Lynch families.33,34 A second disease locus was linked to 3p, and by early 1994 MLH1 germline mutations were noted in Lynch kindreds.35-37 The demonstration of disease causing mutations in PMS2 and MSH6 would follow.38-40 The next 15 years have seen the application of this knowledge to the better understanding of disease incidence and phenotype and to the exploration of clinical issues of detection and management. Especially notable in this time period are the 2 National Cancer Institute-sponsored (NCI) workshops dedicated to the issuance of guidelines (“Bethesda guidelines’’) to identify patients who would benefit from clinical testing for Lynch syndrome.11,12 The Revised Bethesda guidelines (BG) are presented in Table 2.
Mismatch Repair
DNA replication is associated with a finite error rate, including the incorporation of mispaired bases (eg, G with A) and slippage of the DNA strands during replication (resulting in the formation of insertion/ deletion loops). Failure to repair these errors results in point and frameshift mutations, respectively. The DNA MMR apparatus recognizes errors that elude the proofreading function of DNA polymerase. The system is highly conserved from bacteria to humans, and as the system had been previously characterized in single-celled organisms, the link between microsatelite instability (MSI) and deficient mismatch repair function (dMMR) in human cancer was first recognized by microbial geneticists.42,43 The human MMR genes are named after their prokaryotic counterparts. For example, MSH2 is “MutS homologue 2’’ (The “mut’’ refers to the generalized hypermutability noted in bacterial strains with loss of MutS function). The MMR proteins function as heterodimers. The MSH2-MSH6 complex recognizes mispaired bases and insertion/deletion loops. It recruits MLH1-PMS2, which subsequently directs the remainder of the MMR machinery.43 MSH2 and MLH1 are the dominant (obligate) constituents of their respective pairs. In the absence of MSH6, MSH2 can pair with MSH3, and in the absence of PMS2, MLH1 can pair with PMS1.
Revised Bethesda Guidelines: To Guide Clinical Testing of Colorectal Tumors for MSI* (Table 2)
- CRC diagnosed in a patient who is <50 years old
- Presence of synchronous or metachronous CRC or other HNPCC-associated tumors, regardless of age**
- CRC with MSI-H histology diagnosed in a patient <60 years old***
- CRC diagnosed in Z1 first-degree relatives with an HNPCC associated tumor, with one of the cancers being diagnosed at <50 years old
- CRC diagnosed in Z2 first-degree or second-degree relatives with HNPCC-associated tumors, regardless of age
J Natl Cancer Inst. 2004;96:261–268.3
*Fulfillment of only 1 criterion necessary to warrant clinical testing.
**HNPCC-related tumors defined here as colorectal, endometrial, stomach, ovarian, pancreas, ureter and renal pelvis, biliary tract, brain (glioblastoma), skin (sebaceous adenoma and keratoacanthoma), and small bowel.
***Tumor-infiltrating lymphocytes, Crohn-like reaction, mucinous/signet ring differentiation, or medullary growth pattern. CRC indicates colorectal cancer; HNPCC, hereditary nonpolyposis colorectal cancer; MSI, microsatellite instability.
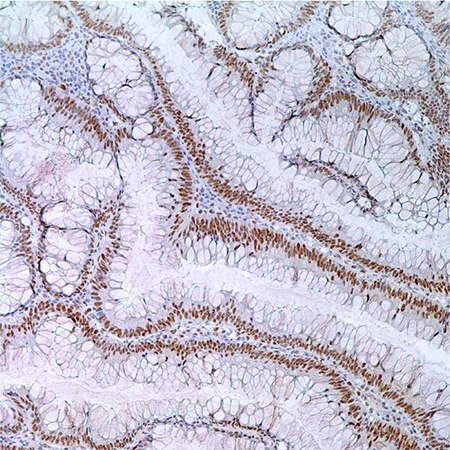
Figure 3. MLH1 (G168-728) in colonic adenoma in 55 year old male; nuclear labeling of adenoma cells.
The MSH6 and PMS2 proteins are unstable in the absence of their respective dominant partner. This was demonstrated by quantitative reverse transcription-polymerase chain reaction and Western blot in a series of dMMR cell lines.50 Also, the majority of Lynch mutations are nonsense or frameshift mutations, leading to truncated (unstable) proteins; missense mutations may destabilize the resulting mRNA or protein or interfere with protein-protein interaction.51 These features of the MMR proteins have clinical diagnostic consequences as relates to the interpretation of MMR immunohistochemistry (IHC) (“Clinical Testing’’).
Microsatellite Instability
Microsatellites are simple repetitive DNA sequences scattered throughout the genome, composed of 1 to 6 base pair units that may repeat up to 100 times. They are inherently hypermutable because of their propensity for strand slippage during DNA replication. The correction of the resulting insertion/deletion loops requires the intact function of the DNA MMR system. With loss of MMR function insertion/deletion loops are not repaired, resulting in the variable expansion or contraction of microsatellites. This phenomenon is referred to as MSI (demonstrated on MSI testing as a typical pattern of bands on a gel or peaks on a sequence analyzer). Genes that contain simple repetitive sequences in critical regions are susceptible to this same phenomenon. The resulting frameshift mutations in MSI-H tumors lead to loss of function in a well-described group of tumor suppressors including TGFbRII, BAX, IGFRII, and, interestingly MSH3 and MSH6.52-55 As such, MSI-H tumors have been described as exhibiting a “mutator phenotype’’.
A 1997 NCI workshop established a reference panel of microsatellites for clinical and research testing and defined diagnostic criteria for the MSI-H, MSI-L, and MSS phenotypes. The core panel consists of 2 mononucleotide repeats (BAT25, BAT26) and 3 dinucleotide repeats (D5S346, D2S123, D17S250). Nineteen “alternative loci’’ are also provided. When analyzing 5 loci, MSI-H is defined as instability at >2 loci and MSI-L as instability at 1 locus. Given uncertainty over the ability of a finite panel to definitively distinguish MSI-L from MSS, tumors exhibiting instability at 0 loci are defined as “MSS or MSI-L.’’ When greater than 5 loci are studied, MSI-H is defined as instability at >30% to 40% of loci tested and MSI-L as instability at <30% to 40% of loci tested.56 A 2002 NCI workshop addended these guidelines, recommending the testing of additional mononucleotide markers in tumors with instability at only dinucleotide loci, as mononucleotide markers are more reliable in the identification of MSI-H tumors.12
Histology
Mecklin et al.41 reported the first systematic histologic evaluation of CRC and adenomas in “CFS’’ (cancer family syndrome) patients in 1986, predating the recognition of MSI and the genetic basis of Lynch syndrome by 7 years. They noted an increased incidence of mucinous carcinomas in cases versus sporadic controls (39% vs. 20%). Poorly differentiated tumors were also more common (24% vs. 12%), but that result failed to reach statistical significance. Although the number of adenomas was similar in the 2 groups, the adenomas of CFS patients tended to contain higher-grade dysplasia and a more substantial villous component, suggesting they were more “advanced’’. Since that time an array of tumor types and histologic features have come to be associated with Lynch syndrome and MSI-H CRC: mucinous, signet-ring cell, and medullary carcinoma, tumor infiltrating and peritumoral lymphocytes, a “Crohn-like’’ inflammatory response, poor differentiation, tumor heterogeneity, and a “pushing’’ tumor border.24,26,41,57-63 Greenson et al.26,62 have further emphasized the presence of any mucinous component, a lack of dirty necrosis, and well-differentiated tumors as correlating with MSI-H status.
By convention, mucinous adenocarcinoma is defined as a tumor composed of >50% mucin and signet-ring cell adenocarcinoma as a tumor containing >50% signet-ring cells. The mucin in mucinous adenocarcinomas is predominantly extracellular, whereas signet-ring cells contain prominent intracytoplasmic mucin, typically displacing the nucleus to one side of the cell.64 Medullary carcinomas are characterized by a constellation of cytoarchitectural features including large cell size, vesicular chromatin and prominent nucleoli, sheet-like to occasionally trabecular growth, and overall circumscription. Tumor infiltrating lymphocytes (TILs) are especially prominent in this tumor type.64
TILs refer to the lymphoid component intimately admixed with the tumor; they have been shown to consist largely of CD3/CD8 co-expressing cytotoxic T-cells [their prominence is speculated to represent: (1) a response to abundant tumor neoantigen formation owing to the “mutator phenotype’’; and (2) a possible basis for the improved prognosis in MSI-H tumors].59 Various methods (and thresholds) for counting TILs have been reported, including evaluation of hematoxylin and eosin or CD3- immunostained slides. One practical method involves scanning the slide for a region with TIL, counting 5 consecutive 40X fields, and calculating the mean number of TIL/high-power field (HPF); studies using this method defined a positive result as >2 TIL/HPF.62 Peritumoral lymphocytes refer to the lymphoid cuff at the leading edge of the tumor, whereas the “Crohn-like’’ reaction is composed of “prominent’’ nodular lymphoid aggregates at the infiltrating edge of the tumor, typically identified at the junction of the muscularis propria and pericolonic adipose tissue. Thresholds for a positive “Crohn-like’’ reaction have included “2 or more large lymphoid aggregates in a section’’, “a single 4X field of at least 3 nodular aggregates of lymphocytes”, “a minimum of 3 lymphoid aggregates per section”, and “at least 4 nodular aggregates in a low power field (4X).”26,61,62,65
Tumors are generally graded as well, moderately, or poorly differentiated. World Health Organization criteria state that tumors with heterogeneous patterns of differentiation should be categorized based on the highest-grade component, with the important caveat that foci of poor differentiation at the leading edge of a tumor (ie, tumor budding, epithelial-mesenchymal transition) are insufficient to classify a tumor as poorly differentiated.64 Tumor heterogeneity refers to the presence of 2 or more distinct growth patterns within a tumor; Alexander and colleagues provide the example of a tumor with mixed mucinous and medullary features.65 A “pushing’’ or “expansile’’ tumor margin is distinguished from an “infiltrative’’ one; this assessment is best made at low power.
Clinical Testing
Clinical testing for dMMR function in CRC has 2 potential endpoints: (1) to identify the Lynch syndrome; or (2) to identify all MSI-H CRC and is associated with 2 fundamental screening strategies: (1) select patients for testing based on clinical and/or pathologic criteria; or (2) test all CRC. In this light, the AC and BG have been evaluated in population-based cohorts for their ability to identify patients with LS. In general, the AC-II achieve sensitivities on the order from 40% to 50%, whereas the revised BG perform at about 90%.13-16,20-23,71 It should be emphasized that these test characteristics are achieved in an idealized setting, with Finnish investigators having access to national cancer registries and other researchers describing the use of “hospital clinical geneticists’’ and “detailed questionnaires’’. Practical barriers to the use of the BG include their complexity and the vagaries in documenting a reliable family cancer history.72
Other limitations of current clinical guidelines include their reduced sensitivity in smaller families and in populations in whom routine screening colonoscopy alters the natural history of the disease. And obviously, these strategies do not target the larger population of MSI-H CRC. The fulfillment of clinical criteria for LS triggers further testing.
If the predictive significance of MSI-H status (ie, therapeutic relevance) can be definitively proven in prospective, randomized controlled clinical trials, then the “test all CRC’’ strategy for an endpoint of MSI-H should become the standard of care. MSI testing is considered the “gold standard’’ for the demonstration of MSI-H status.
As nondirected germline mutation testing for Lynch syndrome is prohibitively expensive (approximately $1000/ gene), MSI testing, and MMR IHC have been evaluated as screening tests.73 Each of these tests has associated advantages and disadvantages. MSI testing involves the PCR-based interrogation of a panel of microsatellite markers in matched tumor and normal tissue (generally adjacent non-neoplastic colon or peripheral blood). It requires microdissection and the services of a molecular diagnostics laboratory. Its inability to suggest a particular gene for mutation analysis is a clear disadvantage, and if further testing is restricted to MSI-H cases, it may also be slightly less sensitive than IHC in the identification of LS due to MSH6 mutation (which may present as MSI-L CRC).74
IHC is readily available in the majority of diagnostic anatomic pathology laboratories. Antibodies to the 4 proteins implicated in LS (through germline mutation of the associated gene) are commercially available. An abnormal result is the complete absence (loss) of nuclear immunoreactivity for one or more of the proteins in the tumor. All 4 proteins are normally expressed in nonneoplastic tissue, and thus stroma, lymphocytes, and nonneoplastic crypts serve as critical internal controls. An antigen retrieval step has been shown to be important; in one study inadequate antigen retrieval was associated with weak staining and poor specificity for LS (whereas high background staining was associated with poor sensitivity).98 Weak or heterogeneous staining patterns are occasionally encountered and may be attributable to variation in tissue fixation or other technical factors. Repeating the IHC may be helpful in these cases. A key advantage to the use of IHC in Lynch screening is its ability to direct gene testing. Loss of MSH2 function (because of deleterious mutation) manifests as absent immunohistochemical (IHC) expression of MSH2 and MSH6, whereas loss of MLH1 function (because of deleterious mutation or promoter hypermethylation) is detectable as absent immunohistochemical expression of MLH1 and PMS2. Isolated absence of MSH6 or PMS2 protein suggests mutation in the respective gene.
The results of MMR IHC and MSI testing have been shown to be largely concordant.15,16,22,23,47,48,76-84 In 1 large study, combined use of MLH1 and MSH2 IHC achieved a sensitivity of 92.3% and a specificity of 100% for the identification of MSI-H tumors (1144 cases tested, 302 of which were MSI-H).77 The addition of MSH6 and PMS2 to the panel should lead to further increased sensitivity, as MSH6 and PMS2 mutation are increasingly recognized as etiologic in a significant minority of Lynch cases.44,45,47,48,79 A potential drawback to IHC is its inability to detect missense mutations that fail to destabilize the resulting mRNA or protein. Owing to this, Burgart has estimated the maximum sensitivity of MMR IHC for detecting LS at 95%.83 In our experience, IHC analysis identifies LS and MSI-H CRC in >90% of cases; MSI testing is similarly sensitive for identifying LS.15,16,21-23,82 In tumors exhibiting absence (loss) of MLH1 protein expression, an additional layer of testing may be used before proceeding to MLH1 mutation analysis: MLH1 promoter methylation testing and/or BRAF mutation analysis. Each of these takes advantage of the unique developmental history of sporadic MSI-H CRC, as discussed previously. In the first instance, methylation-specific PCR is used to determine the methylation status of the MLH1 promoter. Methylated cases are likely sporadic in nature, and thus, MLH1 germline mutation analysis may not be necessary. There is one important caveat. Although the “second-hit’’ in the majority of LS is either loss of heterozygosity or somatic mutation, in rare cases MLH1 promoter hypermethylation has been described.27,28,68,85,86 It is also more expensive than BRAF analysis, as well as technically challenging.
Somatic activating mutations in BRAF, a component of the Ras/Raf/MAP kinase pathway, occur in a wide variety of human tumors, including 15% of CRC.143
They are frequently encountered in sessile serrated adenomas and sporadic MSI-H CRC (around 75%), and are taken as evidence of their evolutionary link.28,66‑69,70 The majority of mutations are accounted for by a point mutation resulting in the substitution of glutamic acid for valine at codon 600 (V600E); in one study of CRC, 60/63 (95%) mutations were V600E.66 Thus, the presence of BRAF V600E in a CRC supports the interpretation that the case is sporadic, and MLH1 germline mutation analysis generally is not pursued. Testing for a single point mutation (rather than sequencing the entire gene) makes BRAF mutation analysis economically attractive ($100).73
Occasionally one may be asked to evaluate adenomas in patients with “possible Lynch syndrome’’, in the setting of a strong family history in which tumor tissue is not available for analysis. MMR IHC is reasonably sensitive and highly specific in this setting.25,87,88 Halvarsson et al.87 reported loss of MMR protein expression in 23/35 (66%) of adenomas from 26 “HNPCC’’ patients (88% of whom had proven germline MMR mutations). Absence of staining was particularly frequent in adenomas >5mm in size (88%). Nineteen of the adenomas demonstrating MMR protein loss contained only typical, low-grade dysplasia. The pattern of absent staining predicted the involved gene in each case.
Absent staining in this clinical setting must be distinguished from that seen in sessile serrated adenomas with superimposed cytologic dysplasia (SSAD) (the generally accepted precursor of sporadic MSI-H CRC).89 The architecture of the lesion and the pattern of loss are important considerations. SSADs are serrated (adenomas in LS are nearly always not), may demonstrate combined loss of MLH1 and PMS2 (not MSH2 and/or MSH6), and, loss of expression is generally considered a late event, frequently corresponding with invasion (whereas loss of expression is often seen in LS adenomas with only low grade dysplasia).
The demonstration of a deleterious germline mutation is the “gold standard’’ for the diagnosis of Lynch syndrome. It allows for relatively inexpensive, directed germline analysis in family members. Following are several points to consider, however. In general, mutation analysis is performed from a peripheral blood sample, including sequence analysis of exons and intron-exon boundaries of the implicated gene. Most reported pathogenic mutations are nonsense or frameshift, resulting in a truncated protein; missense mutations are more difficult to classify.51 Large deletions may account for up to 20% of pathogenic mutations, and these are not detected with routine sequencing. Multiplex ligation-dependent probe amplification (MLPA) provides a quantitative measure of exon dosage and is considered the method of choice for the detection of large deletions.90-93 With hundreds of described mutations in LS, mutation analysis is complex, and the sensitivity of testing, although continuing to improve, is less than 100%. In the setting of a compelling family history, the inability to document an unequivocally pathogenic germline mutation should not exclude a patient from LS-appropriate surveillance. A Lynch syndrome mutational database is maintained by the International Society for Gastrointestinal Hereditary Tumors (formerly the International Collaborative Group on Hereditary Non Polyposis Colorectal Cancer).94
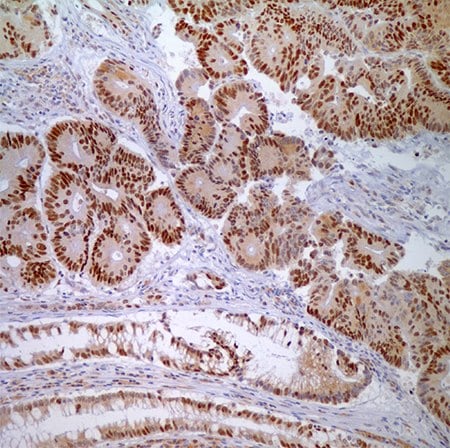
Figure 4. MSH6 (44) - nuclear labeling of adenocarcinoma cells and normal crypt epithelial cells.
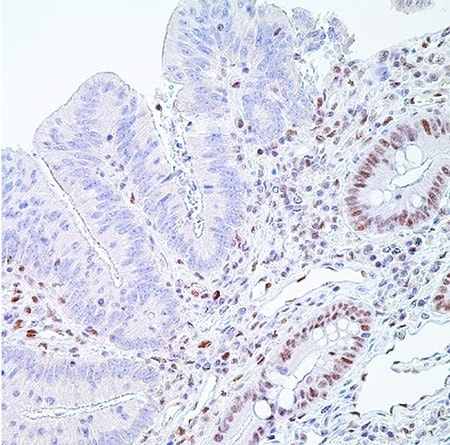
Figure 5. PMS2 (EPR3947†)– Note the normal nuclear labeling of the crypt epithelial cells (and scattered lymphocytes in the lamina propria) to the right and absence of nuclear labeling in the adenocarcinoma cells on the left representing a positive test (MSI-H).
Prognostic Significance of Mismatch Repair Deficiency in Colorectal Cancer
The selectable advantage conferred on tumor cells by the acquisition of MMR deficiency is contentious and may, in part, relate to changes in susceptibility to apoptosis.95,96,97,98 It was noted even at the initial description of MSI, that those patients with MSI-H tumors appeared to have better survival rates than those with MSS tumors.21 Since then, there have been many studies addressing the prognostic significance of MSI-H in CRC, with most of them agreeing with the initial findings,99-102 particularly in young patients, and a few studies failing to identify MSI status as an independent prognostic factor.103,104 In apparent support of the prognostic value of the MSI status is the high prevalence of activated intraepithelial cytotoxic T lymphocytes and increased neoplastic cell apoptosis in MSI-H tumors.99,101 The former characteristic could be attributable to the inherent capacity of MSI-H tumors to produce new immunogenic epitopes (if B2M and HLA class I loci are intact and these novel antigens can be presented), and this could explain why some patients with MSI-H tumors have a particularly favorable clinical outcome following effective anti-tumor immune responses.
The predictive significance of MSI-H has also been assessed in relation to the selection of CRC patients to receive adjuvant chemotherapy. The main correlation that has been shown is the recognition of 5-fluorouracil (5FU)-induced DNA lesions by the intact MMR system105-107 resulting in MMR-deficient cells being more resistant to 5-FU than MMR-proficient cells. Interestingly, when the MMR deficiency was due to MLH1 hypermethylation the cells regained their sensitivity to 5-FU upon MLH1 demethylation.108 However, a clearcut role for MSI-H as a predictor of response to chemotherapy is still controversial.109-111
Although MSI testing can be carried out relatively easily in molecular diagnostic laboratories, there are doubts over the clinical utility of MSI-H as a prognostic marker in sporadic CRC.112,113 An important factor in the failure to reach a consensus is the variation in the type and number of microsatellite markers that have been assessed by different studies. A large meta-analysis 114 with >7000 patients has shown that there is a clear correlation between MSI-H tumors and improved overall survival. However, this study was performed retrospectively and thus could be liable to various confounding factors. It is also possible that differences in the underlying molecular pathways of LS and sporadic MSI-H tumors, as shown by the mutation frequency data in Table 1, also play a part, and thus there may be important differences between LS and sporadic MSI-H tumors that are obscured by combination of their data.
Genetic Counseling and Management of Hereditary Nonpolyposis Colon Cancer
In determining the appropriate time to do genetic testing and begin a surveillance program, age at onset and natural history of colon carcinomas should be considered. The onset of colorectal carcinoma in Lynch syndrome appears to occur 15 to 20 years earlier than in the general population.10 In Mecklin’s series in Finland119, the mean age of colon carcinoma onset was 41 years, with an age range of 19 to 83 years; the gene penetrance was estimated as 89%. In Lynch’s series in Nebraska the mean age at diagnosis was 44.4 years.117,120 The majority of tumors occur by age 60, with the peak in the 40–50 age range. Thus the target of genetic testing and surveillance programs should be the 25–60 age group.8,117 Most patients with HNPCC have a mutation in one of two DNA mismatch repair genes, hMSH2 or hMLH1. More than 90% of CRC patients with hMSH2 or hMLH1 demonstrate high-frequency MSI (MSI-H)121,122; a lot of these tumors have replication errors.144 Initial germline testing is acceptable for those at high risk for carrying a mutation, such as patients who meet the Amsterdam criteria I or the first three criteria of the Bethesda Guidelines.123 Any patient with CRC with a personal or family history of another CRC or endometrial carcinoma, or who was diagnosed before 50 years of age, regardless of personal or family history, meets the modified guidelines.
In large studies12,29 comparing the cost effectiveness of several strategies for genetic testing of patients with HNPCC, it has been shown that germline testing on CRC probands who satisfy the Amsterdam criteria detects the fewest gene carriers and has a low cost, whereas tumor MSI testing of all CRC patients has the highest cost and detects the most gene carriers. Also, it has been shown that germline hMSH2/ hMLH1 testing of patients who meet the Amsterdam criteria and tumor MSI analysis of the remainder of patients, who meet the modified criteria, with subsequent germline testing of those with an MSI-H tumor, is the best strategy in terms of cost effectiveness.122 If a patient meets the Amsterdam criteria I or is suspected to belong to an HNPCC family, the first screening test should be immunohistochemistry for the detection of hMLH1 and hMSH2 proteins.123
Immunohistochemistry is performed using paraffin-embedded tissue blocks, which are counterstained with Mayer hematoxylin. The hMLH1 and hMSH2 proteins stain positively in cell nuclei when expressed. Microsatellite analysis is usually performed with PCR DNA analysis. Several loci are evaluated, such as BAT26, TGF-RII, D2S119, D3S1612, D5S404, D17S261, BAT25, D2S123, D5S346, and D17S250. Tumor DNA strands and normal DNA are compared in neighboring lanes. Tumors are considered MSI-H (high instability) if two or more of the MS markers show instability, MSI-L (low instability) if one marker shows positive instability, and MSS (stable) if no marker shows instability.124 If immunohistochemistry is positive, direct exon-by-exon genomic sequencing should follow for the detection of hMSH2 or hMLH1 mutations. Two of the most commonly occurring mutations are founder mutation 1, a genomic deletion of MLH1 comprising exon 16, and founder mutation 2, an MLH1 exon 6 splice site mutation IVS5-1G»A at 454-1.125
If the test is negative, then MS analysis should follow. If the latter is negative, no more examinations are needed. If it is positive, it should be followed by mutation analysis of MLH1 and MSH2, which is done by sequencing of the coding exons, including the flanking intronic regions and promoter region124-126 (Figure 2). None of the three methods can be used individually for the genetic diagnosis of HNPCC. A combination of these methods and the family history should be used. Southern blot analysis should also be used as a supplemental test to reveal severe rearrangements of the genome. When a germline mutation is identified in affected relatives in an HNPCC family, this will place their lifetime risk for cancer in the range of 80–85%.120 The process of molecular evaluation includes standard and alternative mutation detection techniques (Table 3).
The patients should be instructed about the natural history of HNPCC, with analysis of the advantages of available surveillance and management strategies.6 The patients must realize that a positive result may produce fear and anxiety in the family. Some members of families with HNPCC may already suffer from unresolved grief due to multiple deaths from cancer in their families. These individuals may be at risk of an adverse psychological reaction and may benefit from a psychiatric consultation.120
All these individuals at risk should be informed that the age at cancer onset for HNPCC tends to be early but may be variable; therefore surveillance should continue throughout their lifetimes. Since the evolution of colorectal adenoma to carcinoma is accelerated in HNPCC,115,121 the patients should undergo colonoscopic evaluation every year, in order to reduce the risk of developing a more advanced stage of cancer.119
The Process of Molecular Evaluation (Table 3)
Standard mutation detection techniques
- Single-strand conformational polymorphism
- Denaturing gradient gel electrophoresis analysis
- DNA sequencing
Alternative mutation detection techniques
- Monoallelic expression analysis
- Southern analysis
- Quantitative polymerase chain reaction
Genetic counseling for high-risk HNPCC family members should begin in the midteens. In families with a trend of early mean age at cancer onset, surveillance should begin at age 25. Because of the predominance of right-sided lesions colonoscopy of the entire large bowel is the mainstay of surveillance for HNPCC.7,116 In experienced hands the cecum can be reached in almost 95% of cases, whereas the perforation rate is not more than 0.2%.10 Colonoscopy also offers the ability to take histological samples and remove polyps. Surveillance is performed until age 60, and if there has been no phenotypic expression of the syndrome by then, patients are followed as per the recommendations of the American Cancer Society for colorectal cancer risk in the general population. Endoscopic findings of severe dysplasia of the mucosa or flat adenomas urge for repeat colonoscopy after 6 months. There is an increased incidence of synchronous and metachronous adenomatous polyps, which represent the premalignant lesion in these Lynch syndromes.127 In families with a history of cancer in extracolonic sites, surveillance includes Pap smear, transvaginal ovarian ultrasound, and endometrial aspiration biopsy every 1 to 2 years starting after age 30. For cancers of the urinary system screening should be done every 1 to 2 years with urinalysis, ultrasound, cystoscopy, and urine cytology. For cancers of the stomach and biliary tract esophagogastroduodenoscopy, liver function tests and transabdominal hepatobiliary ultrasound should be performed every 1 to 2 years, beginningat age 30.5
At the time of a newly diagnosed colorectal cancer in a known gene carrier or in a high-risk individual based on family history, a subtotal colectomy with ileorectal anastomosis is the option of choice due to the high risk of metachronous lesions.5,120,128,129 With colorectal cancer risk approaching 80–85%, prophylactic colectomy has been adopted.77 Annual endoscopic examination in the remaining rectum should be instituted. The risk of developing rectal cancer after an abdominal colectomy is estimated to be 12% at 12 years.131 An alternative approach could be a proctocolectomy, especially in the case of an hMSH2 gene carrier, in whom there is a higher risk of rectal cancer than in hMLH1 carriers. Segmental resection is suggested for older patients131 or for patients with co‑morbid conditions, whose life expectancy is already compromised by their preexisting disease.132 In Lynch syndrome II in women, transvaginal ultrasound screening is the preferred method of surveillance. It is a noninvasive, readily available test that allows full examination of the endometrial cavity. Other surveillance methods include endometrial cytologic sampling and endometrial biopsy when indicated. In Lynch syndrome II, women who are prone to develop endometrial cancer and have completed their families or are postmenopausal, total abdominal hysterectomy with bilateral salpingo-oophorectomy should be considered.5,120,127,132,133
Physicians should take time to inform patients of the natural history of this hereditary cancer syndrome, discuss their fears and anxieties about cancer, and advise them about the advantages and disadvantages of genetic testing.
Prognosis
The overall experience of several centers with longterm follow-up of HNPCC patients treated for CRC is encouraging. Several studies report an improved 5-year survival rate among HNPCC patients with colorectal cancer.118,119,134,135 In 1996, Sankila et al. studied 175 patients with HNPCC and compared them in terms of survival with a population of 14,086 patients with sporadic colorectal cancer.136 The diagnosis of the syndrome was based on detection of germline mutations in a mismatch repair gene. The 5-year survival rate for patients with HNPCC was 65%, versus 44% for the sporadic cases. One year later, Watson et al.137 provided additional data on the prognosis of patients with HNPCC. They studied retrospectively a cohort of patients with HNPCC and compared them with a population of patients with sporadic colorectal cancer. They found that over the first 10 years after diagnosis, death rates for HNPCC cases were almost two-thirds the rates for those with sporadic cancer. More recently the Surveillance, Epidemiology, and End Results (SEER) cancer registry has published similar results; age adjusted survival was higher for HNPCC carriers with colorectal cancer than for patients with sporadic colorectal cancer.138,139 Life expectancy was one-third higher for patients with HNPCC and early-stage cancer than for sporadic cases; however, life expectancy decreased with more advanced stages at diagnosis.140
The biological basis for these findings has not been elucidated yet. It has been proposed that the less aggressive behavior of HNPCC cancers may be a paradoxical effect of genomic instability.141 Thus the malignant cell’s essential functions and, especially, their metastatic potential may be inhibited by a very significant mutational load. In addition, gene– gene and gene–environment interactions may play an important role in the natural history of HNPCC. Systematic surveillance and individually designed treatment of affected patients may help to detect cancers at an earlier stage and subsequently further improve the prognosis of the disease.
References
続きを確認するには、ログインするか、新規登録が必要です。
アカウントをお持ちではありませんか?