MOD50
Imprint® DNA Modification Kit
For bisulfite DNA conversion & purification
Synonym(s):
Methylation-Specifc PCR preparation, bisulfite conversion kit, bisulphite modification
About This Item
Recommended Products
General description
Application
Features and Benefits
- Only 50 picograms of DNA or 20 cells are required
- Procedure takes less than 2 hours
- Greater than 99% conversion rate
- Extremely low degradation
- Option of convenient one-step protocol
- Consistent and reproducible Bisulfite Modification
- Can be used with genomic, endonuclease digested, and FFPE DNA
Storage and Stability
Legal Information
Kit Components Also Available Separately
- T3566Clear-view™ Snap-Cap microtubes, size 1.5 mL, naturalSDS
Signal Word
Danger
Hazard Statements
Precautionary Statements
Hazard Classifications
Acute Tox. 4 Oral - Aquatic Chronic 3 - Eye Dam. 1 - Met. Corr. 1 - Skin Corr. 1A
Supplementary Hazards
Storage Class Code
8B - Non-combustible corrosive hazardous materials
Flash Point(F)
Not applicable
Flash Point(C)
Not applicable
Regulatory Listings
Regulatory Listings are mainly provided for chemical products. Only limited information can be provided here for non-chemical products. No entry means none of the components are listed. It is the user’s obligation to ensure the safe and legal use of the product.
PDSCL
Please refer to KIT Component information
PRTR
Please refer to KIT Component information
FSL
Please refer to KIT Component information
ISHL Indicated Name
Please refer to KIT Component information
ISHL Notified Names
Please refer to KIT Component information
Cartagena Act
Please refer to KIT Component information
JAN Code
キットコンポーネントの情報を参照してください
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
Articles
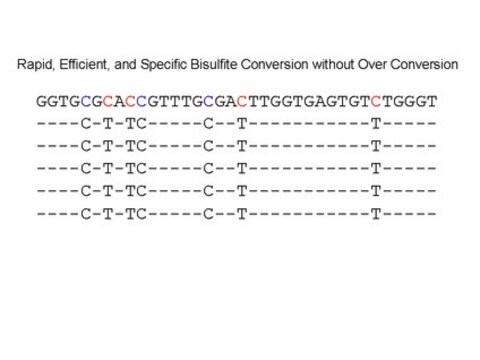
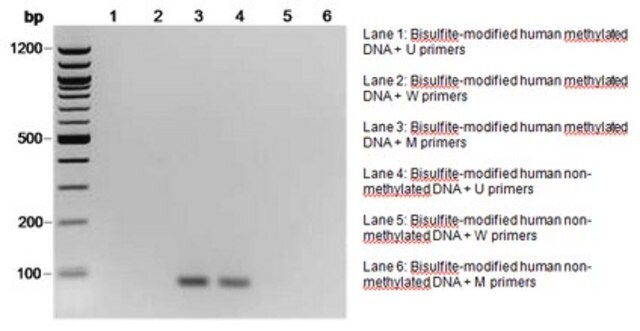
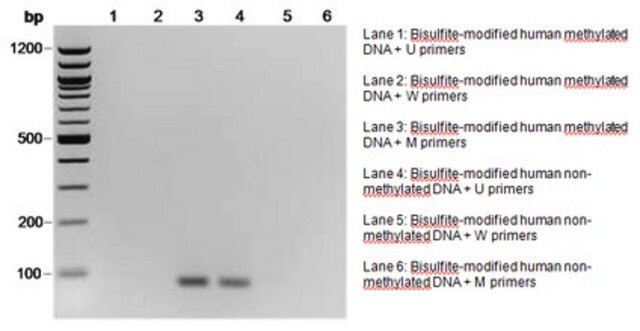
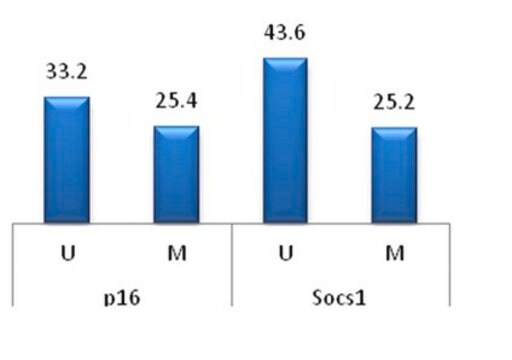
There are several common ways to determine whether a gene contains methylated DNA. Since mammalian methylation occurs at cytosines, researchers take advantage of the fact that methylated cytosine (meC) is stable to bisulfite treatment but unmethylated cytosine is transformed to uracil under the same conditions.
Protocols
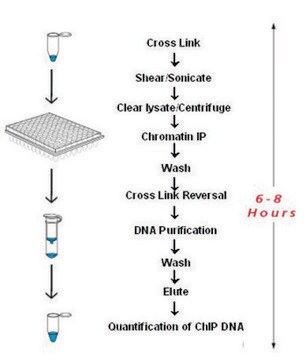
Chromatin Immunoprecipitation qPCR for studying gene regulation across conditions.
Related Content
The Imprint DNA Modification Kit provides the reagents needed for bisulfite conversion and post-modification clean-up of DNA samples in less than 2 hours.
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service