11277081001
Roche
Hexanucleotide Mix
solution, pkg of 100 μL, sufficient for 50 labeling reactions
Sinonimo/i:
nucleotide mix
Autenticatiper visualizzare i prezzi riservati alla tua organizzazione & contrattuali
About This Item
Codice UNSPSC:
41106300
Prodotti consigliati
Stato
solution
Livello qualitativo
impiego
sufficient for 50 labeling reactions
Confezionamento
pkg of 100 μL
Produttore/marchio commerciale
Roche
tecniche
Northern blotting: suitable
Southern blotting: suitable
cDNA synthesis: suitable (first strand)
hybridization: suitable
Colore
colorless
Solubilità
water: miscible
Temperatura di conservazione
−20°C
Descrizione generale
Convenient oligonucleotide mixture for rapid random-primed labeling of DNA with radioactive, digoxigenin- or biotin-labeled nucleotides. In this method, the complementary DNA strand is synthesized by Klenow polymerase using the 3′-OH termini of the random oligonucleotides as primers.
Specificità
Heat inactivation: Stop the reaction by adding 2 μl 0.2 M EDTA (pH 8.0) and/or by heating to 65 °C for 10 minutes.
Applicazioni
Hexanucleotide Mix is a mixture of hexanucleotides of all possible sequences for random-primed DNA labeling.
Labeled DNA probes with high specific activity are used in a variety of hybridization techniques:
Labeled DNA probes with high specific activity are used in a variety of hybridization techniques:
- Screening of gene libraries
- Southern and northern blots
- In situ hybridizations
- RT-PCR
- Generation of cDNA libraries
- Synthesis of first-strand cDNA
- in the determination of vector titer
- Second strand synthesis
Caratteristiche e vantaggi
The product is a 10x concentrated mixture of random hexanucleotides. Statistically, the mix may contain up to 4,096 different hexanucleotides, but these are probably present in differing amounts.
Contents
10x concentrated mixture of hexanucleotides (62.5 A260 units/ml) in reaction buffer [0.5M Tris- HCl, 0.1M MgCl2, 1mM dithioerythritol (DTE), 2mg/ml BSA, pH 7.2 (+20°C)]
Note: The mix is identical to that supplied in vial 5 of the DIG DNA Labeling and Detection Kit and of the DIG DNA Labeling Kit and in vial 6 of the Random Primed DNA Labeling Kit.
Contents
10x concentrated mixture of hexanucleotides (62.5 A260 units/ml) in reaction buffer [0.5M Tris- HCl, 0.1M MgCl2, 1mM dithioerythritol (DTE), 2mg/ml BSA, pH 7.2 (+20°C)]
Note: The mix is identical to that supplied in vial 5 of the DIG DNA Labeling and Detection Kit and of the DIG DNA Labeling Kit and in vial 6 of the Random Primed DNA Labeling Kit.
Qualità
Function tested in the Random Primed DNA Labeling Kit.
Principio
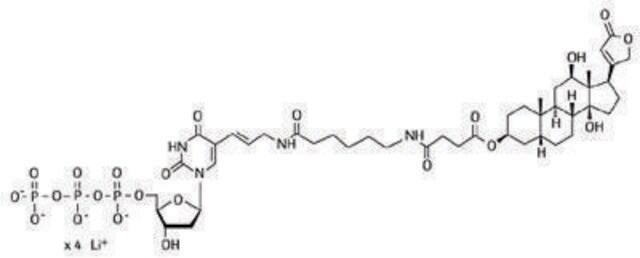
The "random primed" DNA labeling method developed by Feinberg and Vogelstein is based on the hybridization of a mixture of all possible hexanucleotides to the DNA to be labeled. All sequence combinations are represented in the hexanucleotide primer mixture, which leads to binding of primer to the template DNA in a statistical manner. Thus, an equal degree of labeling along the entire length of the template DNA is guaranteed. The complementary strand is synthesized from the 3′-OH termini of the random hexanucleotide primer using Klenow enzyme, labeling grade. Modified deoxyribonucleoside triphosphates ([32P], [35S],[3H], or [125I] digoxigenin- or biotin-labeled) present in the reaction are incorporated into the newly synthesized complementary DNA strand.
Nota sulla preparazione
Assay Time
Standard labeling (radioactive): 50 minutes
Labeling assay with digoxigenin-11-dUTP: 80 minutes
Sample Materials
Synthesis: All 4 bases are used to synthesize this random hexanucleotide mix. In the initial reaction, starter nucleotides are linked to a solid phase support. In subsequent coupling reactions, equimolar amounts of the 4 dNTPs are linked to the starter nucleotides until hexamers are generated. The hexamers are then released from the solid phase support.
Post-synthesis: The oligonucleotides are HPLC purified, desalted, and 5′-phosphorylated.
Standard labeling (radioactive): 50 minutes
Labeling assay with digoxigenin-11-dUTP: 80 minutes
Sample Materials
- DNA fragments
- Linearized plasmid DNA
- λDNA
Synthesis: All 4 bases are used to synthesize this random hexanucleotide mix. In the initial reaction, starter nucleotides are linked to a solid phase support. In subsequent coupling reactions, equimolar amounts of the 4 dNTPs are linked to the starter nucleotides until hexamers are generated. The hexamers are then released from the solid phase support.
Post-synthesis: The oligonucleotides are HPLC purified, desalted, and 5′-phosphorylated.
Risultati analitici
Absorption: 62.5 A260 units correspond to 2.5 mg/ml of hexanucleotides.
Altre note
For life science research only. Not for use in diagnostic procedures.
Codice della classe di stoccaggio
12 - Non Combustible Liquids
Classe di pericolosità dell'acqua (WGK)
WGK 1
Punto d’infiammabilità (°F)
No data available
Punto d’infiammabilità (°C)
No data available
Scegli una delle versioni più recenti:
Possiedi già questo prodotto?
I documenti relativi ai prodotti acquistati recentemente sono disponibili nell’Archivio dei documenti.
I clienti hanno visto anche
Chen Ling et al.
Journal of visualized experiments : JoVE, (49)(49), doi:10-doi:10 (2011-03-30)
Recombinant vectors based on a non-pathogenic human parvovirus, the adeno-associated virus 2 (AAV2) have been developed, and are currently in use in a number of gene therapy clinical trials. More recently, a number of additional AAV serotypes have also been
Alterations in ribosome biogenesis cause specific defects in C. elegans hermaphrodite gonadogenesis.
Roumen Voutev et al.
Developmental biology, 298(1), 45-58 (2006-08-01)
Ribosome biogenesis is a cell-essential process that influences cell growth, proliferation, and differentiation. How ribosome biogenesis impacts development, however, is poorly understood. Here, we establish a link between ribosome biogenesis and gonadogenesis in Caenorhabditis elegans that affects germline proliferation and
Elise Alspach et al.
Bio-protocol, 5(10) (2015-05-20)
Immunoprecipitation and subsequent isolation of nucleic acids allows for the investigation of protein:nucleic acid interactions. RNA-binding protein immunoprecipitation (RIP) is used for the analysis of protein interactions with mRNA. Combining RIP with quantitative real-time PCR (qRT-PCR) further enhances the RIP
Andrew L Goodman et al.
Nature protocols, 6(12), 1969-1980 (2011-11-19)
Insertion sequencing (INSeq) is a method for determining the insertion site and relative abundance of large numbers of transposon mutants in a mixed population of isogenic mutants of a sequenced microbial species. INSeq is based on a modified mariner transposon
Dilip Kumar et al.
Journal of immunology (Baltimore, Md. : 1950), 182(2), 1011-1020 (2009-01-07)
The MAPKs ERK, JNK, and p38 control diverse aspects of the immune response, including regulation of cytotoxin biology in NK cells and CTL. The chemokine CCL5 is coreleased with the cytotoxins, perforin, the granzymes, and granulysin, during the lethal hit
Il team dei nostri ricercatori vanta grande esperienza in tutte le aree della ricerca quali Life Science, scienza dei materiali, sintesi chimica, cromatografia, discipline analitiche, ecc..
Contatta l'Assistenza Tecnica.