T3411
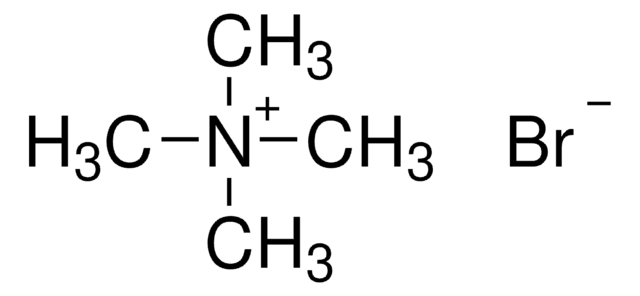
Tetramethylammonium chloride solution
for molecular biology
Sinónimos:
N,N,N-Trimethylmethanaminium chloride
Iniciar sesiónpara Ver la Fijación de precios por contrato y de la organización
About This Item
Número de CAS:
Número MDL:
Código UNSPSC:
12352107
ID de la sustancia en PubChem:
NACRES:
NA.31
Productos recomendados
grado
for molecular biology
Nivel de calidad
concentración
5 M
actividad extraña
DNase, RNase, none detected
cadena SMILES
[Cl-].C[N+](C)(C)C
InChI
1S/C4H12N.ClH/c1-5(2,3)4;/h1-4H3;1H/q+1;/p-1
Clave InChI
OKIZCWYLBDKLSU-UHFFFAOYSA-M
Categorías relacionadas
Descripción general
Tetramethylammonium binds AT-rich DNA polymers while concomitantly abolishing the preferential melting of AT versus GC base pairs. It is supplied as a 0.2 μm filtered solution in 18 megohm water.
Aplicación
Tetramethylammonium chloride solution (TMAC) has been used:
- in the preparation of hybridization cocktail for array hybridization and scanning
- in next-generation sequencing (NGS), and genome-wide unbiased identification of double-stranded breaks enabled by sequencing (GUIDE-seq) library preparation
- in the preparation of TMAC buffer and bead hybridization mixture for hybridization and detection
Elija entre una de las versiones más recientes:
¿Ya tiene este producto?
Encuentre la documentación para los productos que ha comprado recientemente en la Biblioteca de documentos.
Los clientes también vieron
W B Melchior et al.
Proceedings of the National Academy of Sciences of the United States of America, 70(2), 298-302 (1973-02-01)
Several small alkylammonium ions can eliminate, or even reverse, the usual dependence of the DNA transition temperature on base composition. For example, in 3 M tetramethylammonium chloride, or 2.4 M tetraethylammonium chloride, DNAs of different base compositions all melt at
Nikolay L Malinin et al.
Nature protocols, 16(12), 5592-5615 (2021-11-14)
Genome-wide unbiased identification of double-stranded breaks enabled by sequencing (GUIDE-seq) is a sensitive, unbiased, genome-wide method for defining the activity of genome-editing nucleases in living cells. GUIDE-seq is based on the principle of efficient integration of an end-protected double-stranded oligodeoxynucleotide
Hybridization of genomic DNA to oligonucleotide probes in the presence of tetramethylammonium chloride.
A G DiLella et al.
Methods in enzymology, 152, 447-451 (1987-01-01)
Xiao-Yong Li et al.
Methods in molecular biology (Clifton, N.J.), 809, 3-26 (2011-11-25)
Immunoprecipitation of cross-linked chromatin in combination with microarrays (ChIP-chip) or ultra high-throughput sequencing (ChIP-seq) is widely used to map genome-wide in vivo transcription factor binding. Both methods employ initial steps of in vivo cross-linking, chromatin isolation, DNA fragmentation, and immunoprecipitation.
Markus Welcker et al.
Genes & development, 27(23), 2531-2536 (2013-12-04)
The Fbw7 tumor suppressor targets a broad network of proteins for ubiquitylation. Here we show critical functions for Fbw7 dimerization in regulating the specificity and robustness of degradation. Dimerization enables Fbw7 to target substrates through concerted binding to two suboptimal
Nuestro equipo de científicos tiene experiencia en todas las áreas de investigación: Ciencias de la vida, Ciencia de los materiales, Síntesis química, Cromatografía, Analítica y muchas otras.
Póngase en contacto con el Servicio técnico