KMM-101NV
KOD One™ PCR Master Mix
Ready-to-use 2X hot-start PCR master mix with a modified KOD DNA polymerase optimized for ultra-fast and convenient high-fidelity PCR
Sinónimos:
Fast Hot-start PCR, High-fidelity DNA polymerase, KOD One™ PCR Ready Mix
About This Item
Productos recomendados
origen biológico
Escherichia coli (carrying the cloned UKOD DNA polymerase gene)
Nivel de calidad
Formulario
liquid
uso
sufficient for 25 reactions
sufficient for 500 reactions
Características
Fast PCR
High Fidelity PCR
dNTPs included
hotstart
técnicas
PCR: suitable
entrada
crude DNA
Condiciones de envío
dry ice
temp. de almacenamiento
−20°C
Descripción general
This hot start KOD polymerase is antibody-inactivated with two types of antibodies that prevent the 5′→3′ polymerase activity and 3′→5′ exonuclease activity. Upon heat-activation, the 3′→5′ proof-reading ability of the enzyme generates blunt-end PCR products.
Aplicación
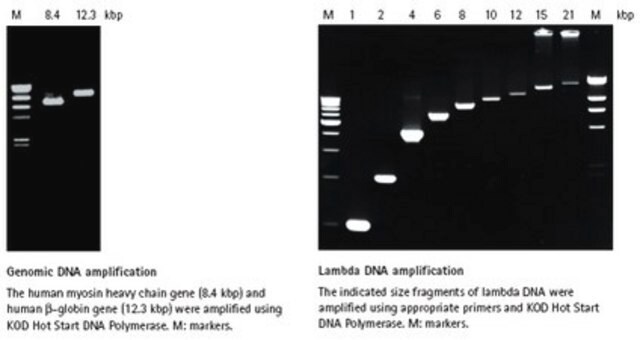
- Genomic DNA amplification
- cDNA amplification
- Direct PCR
- Colony PCR
- Amplification of NGS libraries
- Site direct gene mutation
Compatible Sample Types: Serum, Plasma, Cell, Plant, soil extraction, DNA, nail, hair etc.
Sample Volume Needed: about 50ng DNA
Características y beneficios
- Fast: Amplifies the targets using the following very short conditions:
o 1~ 10 kb: 5 sec/ kb
o 10 kb~: 10 sec/ kb
- Ready-to-use: 2X premixed solution of UKOD DNA polymerase, buffer, dNTPs, and MgCl2.
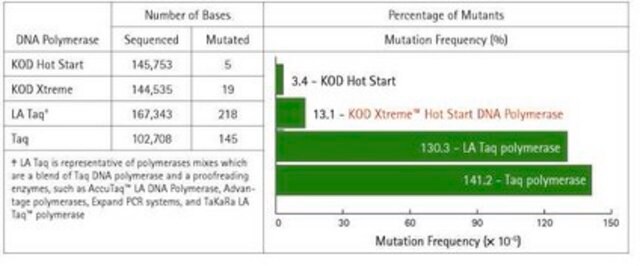
- High Fidelity: Approximately 80-fold higher fidelity than conventional Taq DNA polymerase
- High Efficiency: Enables high throughput PCR and ultra-fast PCR cycling conditions with an extension time of as quick as 5 sec/kb.
- Crude Sample PCR: Effective for direct PCR amplification of crude samples such as biological samples, food samples, soil extract, etc.
- High flexibility: Amplifies templates containing uracils (dU). Also, amplifies fragments using degenerate primers with inosines (dI) and uracils (dU)
Envase
KMM-101NVS - 0.25ML is sufficient for 25 reactions at 20 μL each or for 10 reactions at 50 μL each.
Otras notas
Información legal
Palabra de señalización
Warning
Frases de peligro
Consejos de prudencia
Clasificaciones de peligro
Met. Corr. 1
Código de clase de almacenamiento
8B - Non-combustible corrosive hazardous materials
Clase de riesgo para el agua (WGK)
WGK 2
Punto de inflamabilidad (°F)
Not applicable
Punto de inflamabilidad (°C)
Not applicable
Listados normativos
Los listados normativos se proporcionan para los productos químicos principalmente. Para los productos no químicos sólo se puede proporcionar información limitada. Si no hay ninguna entrada, significa que ninguno de los componentes está en la lista. Es obligación del usuario garantizar el uso seguro y legal del producto.
EU REACH Annex XIV (Authorisation List)
Elija entre una de las versiones más recientes:
Certificados de análisis (COA)
¿No ve la versión correcta?
Si necesita una versión concreta, puede buscar un certificado específico por el número de lote.
¿Ya tiene este producto?
Encuentre la documentación para los productos que ha comprado recientemente en la Biblioteca de documentos.
Contenido relacionado
KOD One™ PCR Master Mix overview for ultra-fast PCR with high specificity, fidelity, and yield
Global Trade Item Number
| Número de referencia del producto (SKU) | GTIN |
|---|---|
| KMM-101NV-5X1ML | 4061841805342 |
| KMM-101NVS-0.25ML | 4061841805359 |
Nuestro equipo de científicos tiene experiencia en todas las áreas de investigación: Ciencias de la vida, Ciencia de los materiales, Síntesis química, Cromatografía, Analítica y muchas otras.
Póngase en contacto con el Servicio técnico