CRISPR31
CRISPR Non-Target Negative Control Plasmid for Bacteria
About This Item
Produits recommandés
Forme
liquid
Conditionnement
vial of 50 μL
Concentration
20 ng/μL in TE buffer; DNA (1μg of purified plasmid DNA)
Technique(s)
microbiological culture: suitable
Application(s)
CRISPR
genome editing
Promoteur
Promoter activity: constitutive
Conditions d'expédition
dry ice
Température de stockage
−20°C
Description générale
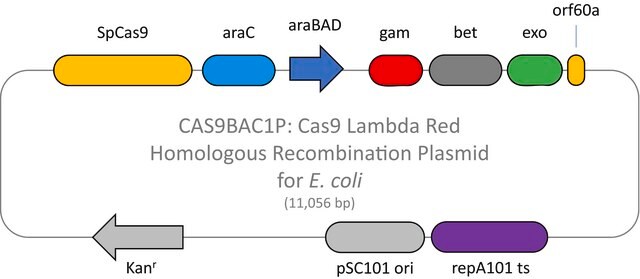
Here we present a novel dual-vector CRISPR/Cas-mediated λ-Red system for improved recombineering in E. coli. Our system is shown to facilitate homology-directed repair of DSBs created by Cas9 endonuclease, enabling genetic alterations through chromosomal integration of a donor DNA.
This plasmid is to be used in combination with the Cas9 Lambda Red homologous recombination plasmid for E. coli (CAS9BAC1P) as the negative control for your custom gene editing experiment. The custom gRNA (CRISPRBACD) can be designed and ordered through https://www.sigmaaldrich.com/pc/ui/genomics-home/customcrispr
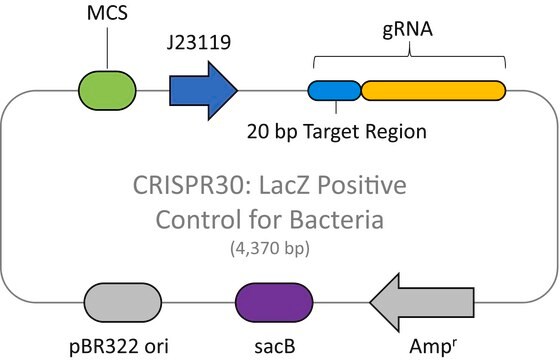
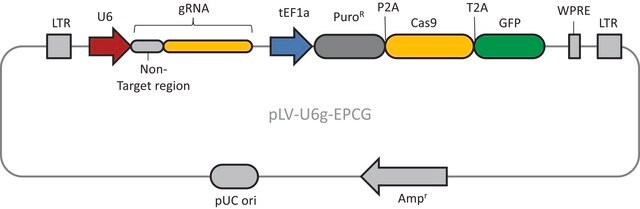
The CRISPR Non-Target Negative Control Plasmid for Bacteria (CRISPR31) contains a non-targeting spacer expressed constitutively from a J23119 promoter, a ampicillin resistance marker, a pBR322 origin of replication, and a sacB gene from Bacillus subtilis for counter-selection-based curing.
Application
- HR-mediated recombineering for mutation or SNP analysis
- Creation of HR-mediated knock-in cell lines with promoters, fusion tags, or reporters integrated into endogenous genes
- Creation of gene knockouts in E. coli cell lines
Strain Optimization
Caractéristiques et avantages
Markerless: does not require antibiotic resistance marker insertion
Scarless: no scar sequences from marker excision which often cause off-target recombination
Multiplexing: multiple custom gRNA sequences can be used at a time
Principe
Informations légales
Code de la classe de stockage
12 - Non Combustible Liquids
Classe de danger pour l'eau (WGK)
WGK 2
Point d'éclair (°F)
Not applicable
Point d'éclair (°C)
Not applicable
Certificats d'analyse (COA)
Recherchez un Certificats d'analyse (COA) en saisissant le numéro de lot du produit. Les numéros de lot figurent sur l'étiquette du produit après les mots "Lot" ou "Batch".
Déjà en possession de ce produit ?
Retrouvez la documentation relative aux produits que vous avez récemment achetés dans la Bibliothèque de documents.
Articles
In this article, we present an application of our novel E. coli CRISPR/Cas-mediated Lambda-Red (λ-Red) homologous recombination (HR) vector system, which facilitates gene editing through the homology-directed repair (HDR) of double-stranded DNA breaks (DSBs) created by Cas9 endonuclease, using either ssDNA or dsDNA as an editing template.
Notre équipe de scientifiques dispose d'une expérience dans tous les secteurs de la recherche, notamment en sciences de la vie, science des matériaux, synthèse chimique, chromatographie, analyse et dans de nombreux autres domaines..
Contacter notre Service technique