CRISPR30
CRISPR LacZ Positive Control plasmid for Bacteria
About This Item
Productos recomendados
formulario
liquid
envase
vial of 50 μL
concentración
20 ng/μL in TE buffer; DNA (1μg of purified plasmid DNA)
técnicas
microbiological culture: suitable
aplicaciones
CRISPR
genome editing
Promotor
Promoter activity: constitutive
Condiciones de envío
dry ice
temp. de almacenamiento
−20°C
Descripción general
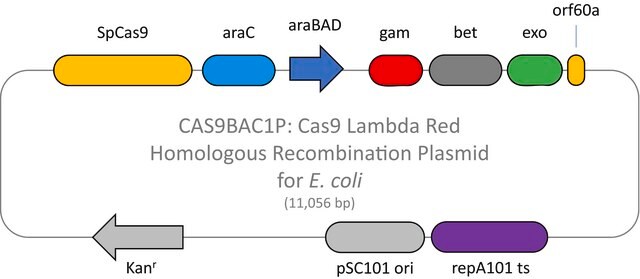
Here we present a novel dual-vector CRISPR/Cas-mediated λ-Red system for improved recombineering in E. coli. Our system is shown to facilitate homology-directed repair of DSBs created by Cas9 endonuclease, enabling genetic alterations through chromosomal integration of a donor DNA.
This plasmid is to be used in combination with the Cas9 Lambda Red homologous recombination plasmid for E. coli (CAS9BAC1P) as the positive control for your custom gene editing experiment. The custom gRNA (CRISPRBACD) can be designed and ordered through https://www.sigmaaldrich.com/pc/ui/genomics-home/customcrispr
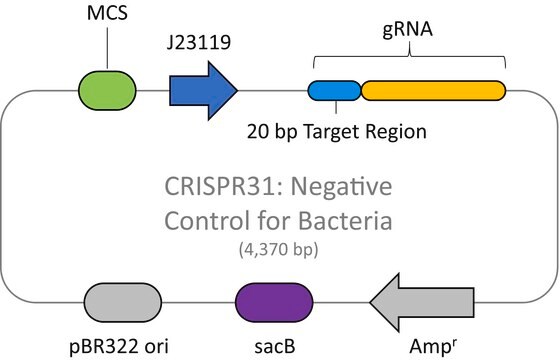
The CRISPR LacZ Positive Control Plasmid for Bacteria (CRISPR30) contains a gRNA spacer targeting the lacZ gene in wild-type E. coli expressed constitutively from a J23119 promoter, a ampicillin resistance marker, a pBR322 origin of replication, and a sacB gene from Bacillus subtilis for counter-selection-based curing.
Aplicación
- HR-mediated recombineering for mutation or SNP analysis
- Creation of HR-mediated knock-in cell lines with promoters, fusion tags, or reporters integrated into endogenous genes
- Creation of gene knockouts in E. coli cell lines
Strain Optimization
Características y beneficios
Markerless: does not require antibiotic resistance marker insertion
Scarless: no scar sequences from marker excision which often cause off-target recombination
Multiplexing: multiple custom gRNA sequences can be used at a time
Principio
Información legal
Producto relacionado
Código de clase de almacenamiento
12 - Non Combustible Liquids
Clase de riesgo para el agua (WGK)
WGK 2
Punto de inflamabilidad (°F)
Not applicable
Punto de inflamabilidad (°C)
Not applicable
Certificados de análisis (COA)
Busque Certificados de análisis (COA) introduciendo el número de lote del producto. Los números de lote se encuentran en la etiqueta del producto después de las palabras «Lot» o «Batch»
¿Ya tiene este producto?
Encuentre la documentación para los productos que ha comprado recientemente en la Biblioteca de documentos.
Artículos
In this article, we present an application of our novel E. coli CRISPR/Cas-mediated Lambda-Red (λ-Red) homologous recombination (HR) vector system, which facilitates gene editing through the homology-directed repair (HDR) of double-stranded DNA breaks (DSBs) created by Cas9 endonuclease, using either ssDNA or dsDNA as an editing template.
Nuestro equipo de científicos tiene experiencia en todas las áreas de investigación: Ciencias de la vida, Ciencia de los materiales, Síntesis química, Cromatografía, Analítica y muchas otras.
Póngase en contacto con el Servicio técnico