Wichtige Dokumente
CRISPR30
CRISPR LacZ Positive Control plasmid for Bacteria
About This Item
Empfohlene Produkte
Form
liquid
Verpackung
vial of 50 μL
Konzentration
20 ng/μL in TE buffer; DNA (1μg of purified plasmid DNA)
Methode(n)
microbiological culture: suitable
Anwendung(en)
CRISPR
genome editing
Promotor
Promoter activity: constitutive
Versandbedingung
dry ice
Lagertemp.
−20°C
Allgemeine Beschreibung
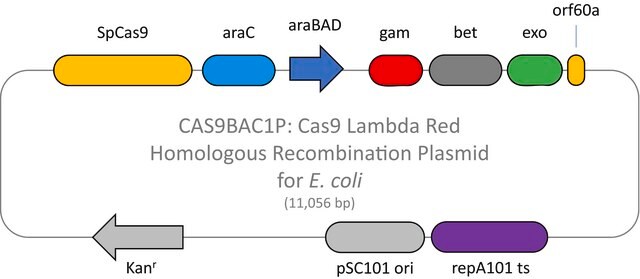
Here we present a novel dual-vector CRISPR/Cas-mediated λ-Red system for improved recombineering in E. coli. Our system is shown to facilitate homology-directed repair of DSBs created by Cas9 endonuclease, enabling genetic alterations through chromosomal integration of a donor DNA.
This plasmid is to be used in combination with the Cas9 Lambda Red homologous recombination plasmid for E. coli (CAS9BAC1P) as the positive control for your custom gene editing experiment. The custom gRNA (CRISPRBACD) can be designed and ordered through https://www.sigmaaldrich.com/pc/ui/genomics-home/customcrispr
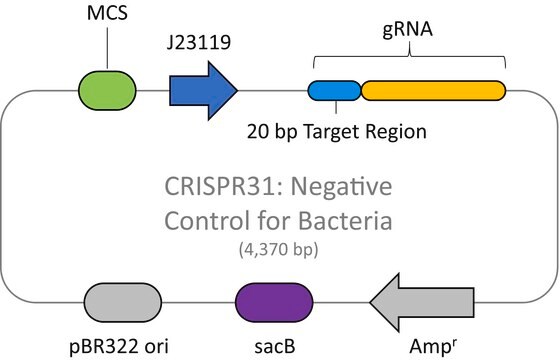
The CRISPR LacZ Positive Control Plasmid for Bacteria (CRISPR30) contains a gRNA spacer targeting the lacZ gene in wild-type E. coli expressed constitutively from a J23119 promoter, a ampicillin resistance marker, a pBR322 origin of replication, and a sacB gene from Bacillus subtilis for counter-selection-based curing.
Anwendung
- HR-mediated recombineering for mutation or SNP analysis
- Creation of HR-mediated knock-in cell lines with promoters, fusion tags, or reporters integrated into endogenous genes
- Creation of gene knockouts in E. coli cell lines
Strain Optimization
Leistungsmerkmale und Vorteile
Markerless: does not require antibiotic resistance marker insertion
Scarless: no scar sequences from marker excision which often cause off-target recombination
Multiplexing: multiple custom gRNA sequences can be used at a time
Prinzip
Rechtliche Hinweise
Ähnliches Produkt
Lagerklassenschlüssel
12 - Non Combustible Liquids
WGK
WGK 2
Flammpunkt (°F)
Not applicable
Flammpunkt (°C)
Not applicable
Hier finden Sie alle aktuellen Versionen:
Analysenzertifikate (COA)
Die passende Version wird nicht angezeigt?
Wenn Sie eine bestimmte Version benötigen, können Sie anhand der Lot- oder Chargennummer nach einem spezifischen Zertifikat suchen.
Besitzen Sie dieses Produkt bereits?
In der Dokumentenbibliothek finden Sie die Dokumentation zu den Produkten, die Sie kürzlich erworben haben.
Artikel
In this article, we present an application of our novel E. coli CRISPR/Cas-mediated Lambda-Red (λ-Red) homologous recombination (HR) vector system, which facilitates gene editing through the homology-directed repair (HDR) of double-stranded DNA breaks (DSBs) created by Cas9 endonuclease, using either ssDNA or dsDNA as an editing template.
Unser Team von Wissenschaftlern verfügt über Erfahrung in allen Forschungsbereichen einschließlich Life Science, Materialwissenschaften, chemischer Synthese, Chromatographie, Analytik und vielen mehr..
Setzen Sie sich mit dem technischen Dienst in Verbindung.