SAB4200625
Anti-Methyl-Histone H3 (Me-Lys9)(H3K9me1) antibody, Mouse monoclonal
clone 7E7-H12, purified from hybridoma cell culture
Synonym(s):
9430068D06RIK, H3.3A, H3.3B, H3F3, H3F3A, H3F3A/H3F3B, H3F3B, HISTONE 3B, LOC100045490, RP11−396C23.1, wu:fb58e10, zgc:56193, zgc:86731
About This Item
Recommended Products
biological source
mouse
Quality Level
conjugate
unconjugated
antibody form
purified immunoglobulin
antibody product type
primary antibodies
clone
7E7-H12, monoclonal
form
buffered aqueous solution
mol wt
antigen ~17 kDa
species reactivity
human
concentration
~1 mg/mL
technique(s)
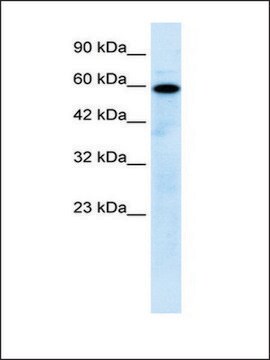
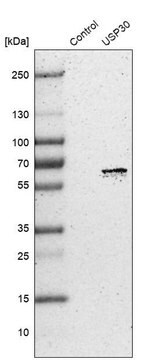
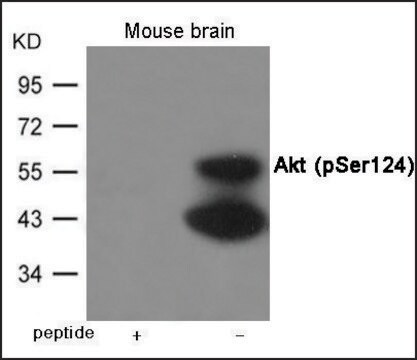
immunoblotting: 2.5-5 μg/mL using human HeLa cells.
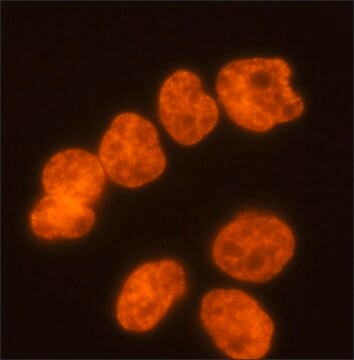
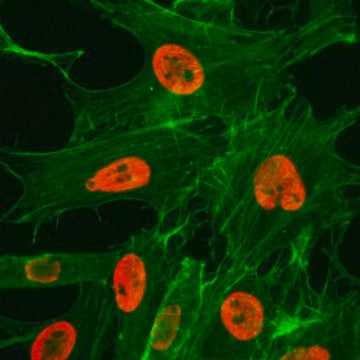
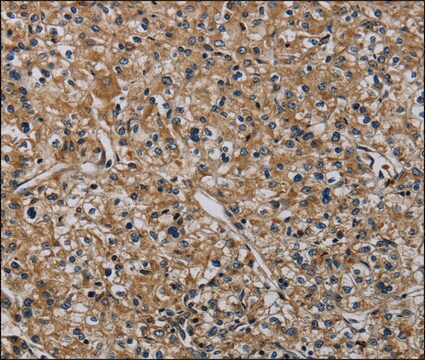
immunocytochemistry: 2.5-5 μg/mL using human HeLa cells.
isotype
IgG1
shipped in
dry ice
storage temp.
−20°C
target post-translational modification
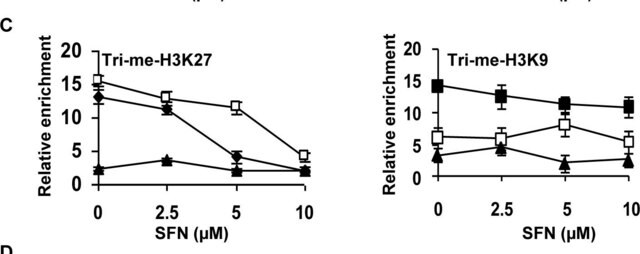
monomethylation (Lys9)
Gene Information
human ... H3C1(8350)
General description
Immunogen
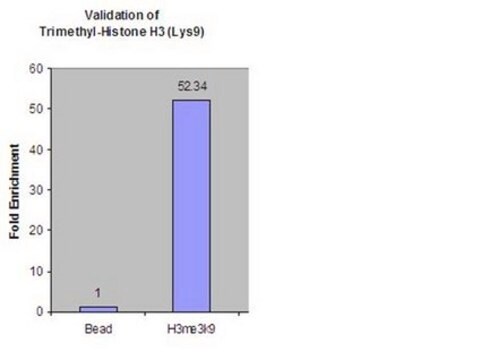
Application
Biochem/physiol Actions
Physical form
Disclaimer
Not finding the right product?
Try our Product Selector Tool.
Storage Class Code
10 - Combustible liquids
WGK
WGK 1
Flash Point(F)
Not applicable
Flash Point(C)
Not applicable
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service