WTA2
Complete Whole Transcriptome Amplification Kit
DNA polymerase included, Complete Kit with optimized enzyme to amplify total RNA in <4 hours, no 3′ bias
Synonym(s):
transcriptome amplification kit
Select a Size
Select a Size
About This Item
Recommended Products
General description
Application
- To establish a protocol for the simultaneous analysis of DNA and RNA viruses present in pig feces using process controlled deep sequencing.
- Reverse transcription and cDNA amplification
- For the synthesis and amplification of cDNA library using Genomic RNA released from immunocaptured PPV particles
- Nucleic Acid Preparation and Deep Sequencing (The extracted nucleic acids were randomly primed for cDNA synthesis)
- qPCR
- microarray analysis
- cloning
Features and Benefits
- Achieve up to 10,000x amplification in less than 4 hours with less than 30 minutes of "hands on" time required
- Only 20 pg of total RNA template is required to amplify suitable cDNA for microarray profiling
- Contains all needed components for cDNA amplification
- Achieve linear amplification of expressed genes and exons without 3′ or 5′ bias
- Effectively amplifies single cell or low input RNA, including mRNA and total RNA from any animal, plant, or microorganism
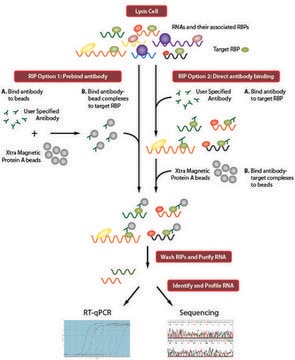
Principle
related product
Storage Class
10 - Combustible liquids
Choose from one of the most recent versions:
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
Articles
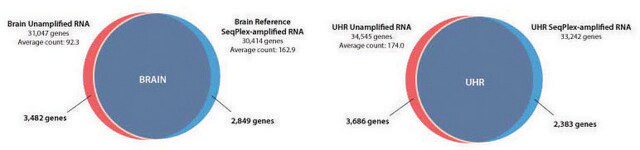
Transplex Whole Transcriptome Amplification (WTA2) exponentially amplifies RNA producing a double-stranded cDNA library while precisely maintaining differential levels of individual transcripts in test and reference samples.
The efficacy of amplification of small quantities of total RNA with the Complete Whole Transcriptome Amplification Kit (WTA2) was examined in this study.
Epigenetic modifications are thought to occur through two key interconnected processes—DNA methylation and the covalent modification of histones.
Protocols
Amplification products generated by the TransPlex® WTA and Complete WTA2 kits are suitable for microarray target for expression analyses, and can be incorporated into existing Illumina workflows.
TransPlex WTA amplification product is suitable as microarray target for expression analysis on the Affymetrix platform, and can be incorporated into existing Affymetrix workflows
Amplification products generated by the TransPlex® WTA and Complete WTA2 kits are suitable for microarray target for expression analyses, and can be readily integrated into existing NimbleGen workflows.
Related Content
Transplex Whole Transcriptome Amplification FAQs on topics including whole transcriptome steps, RNA source, including archival fixed tissue, library purification, quantitation of the product and downstream applications
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service